In the mid-20th century, the discovery of penicillin was hailed as a “miracle cure.” It transformed once-fatal infections into manageable ailments, extending the human lifespan by decades. However, the miracle is fading. We are currently facing a global health crisis known as Antimicrobial Resistance (AMR).
Bacterial resistance to drugs is a condition in which bacteria that were previously susceptible to antibiotics develop the ability to survive and multiply despite the presence of those same drugs. These “superbugs” render standard treatments ineffective, leading to prolonged illnesses, higher medical costs, and an increased risk of death.
Antibiotic Resistance: Hospital vs. Community
While antibiotic resistance can occur anywhere, it is significantly more prevalent in hospital-acquired infections (nosocomial infections) than in community-acquired ones.
Why Hospitals are Hotbeds for Resistance
Selective Pressure: Hospitals use vast quantities of antibiotics. This environment kills off susceptible bacteria but allows resistant strains to survive and thrive—a process known as “selection.”
Vulnerable Populations: Patients in hospitals often have weakened immune systems or invasive devices (like catheters or ventilators), providing easy entry points for resistant bacteria.
Transmission Hubs: Despite strict hygiene protocols, the high density of infected individuals and frequent contact with healthcare workers facilitate the spread of hardy bacterial strains.
These hospital strains are particularly dangerous because they often exhibit multidrug resistance (MDR), meaning they are immune to several different classes of antibiotics simultaneously.
Common “Superbug” Strains
Several specific bacteria have become notorious for their drug-resistant capabilities:
Staphylococcus aureus (MRSA): Often resistant to methicillin and other common penicillins.
Escherichia coli (E. coli): A Gram-negative enteric bacterium increasingly resistant to carbapenems, our “last-resort” antibiotics.
Pseudomonas aeruginosa: A resilient pathogen known for causing severe lung and blood infections, often showing resistance to almost all available treatments.
6 Key Mechanisms of Antibiotic Resistance
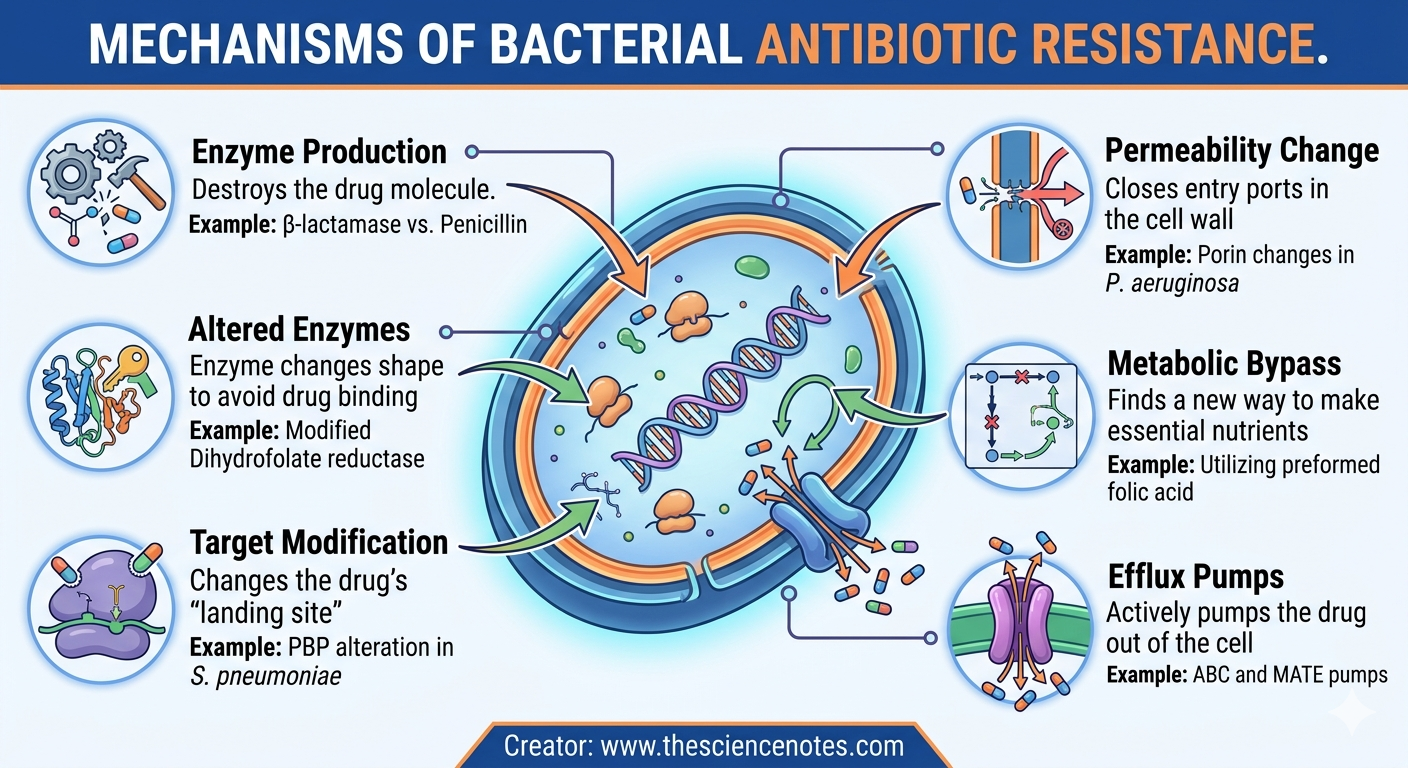
Bacteria are biological masters of adaptation. They employ several sophisticated strategies to neutralize the threat of antibiotics. Understanding these mechanisms is the first step in developing new countermeasures.
1. Production of Inactivating Enzymes
The most common way bacteria fight back is by producing enzymes that physically dismantle the antibiotic molecule before it can do harm.
beta-lactamases: Penicillin-resistant staphylococci produce these enzymes to split the $\beta$-lactam ring (the functional core) of penicillins and cephalosporins.
Carbapenems: History, Structure, Mechanism of Action, resistance and Antimicrobial Activity
Modifying Enzymes: Gram-negative bacteria often carry plasmids that encode adenylating, phosphorylating, or acetylating enzymes. These chemically “tag” aminoglycoside antibiotics, preventing them from binding to their targets.
Aminoglycosides: History, Structure, Antimicrobial Activity, Mechanism of Action, and Resistance
2. Production of Altered Enzymes
Sometimes, bacteria don’t destroy the drug; instead, they change the “lock” so the “key” (the drug) no longer fits.
Example: In trimethoprim-resistant bacteria, the enzyme dihydrofolate reductase is slightly modified. The modified version can still perform its essential metabolic functions but is no longer inhibited by the drug.
3. Synthesis of Modified Targets
Antibiotics work by binding to specific structures within the bacteria, such as ribosomes or cell wall proteins. Bacteria can change these targets to evade detection.
Ribosomal Modification: A methylated 23S ribosomal RNA can cause resistance to erythromycin. Similarly, a mutation in the 50S ribosomal subunit can render streptomycin useless.
PBP Alteration: In S. pneumoniae, resistance to penicillin is often caused by the loss or alteration of Penicillin-Binding Proteins (PBPs), meaning the drug has nothing to latch onto.
4. Alteration of Cell Wall Permeability
If the drug can’t get into the cell, it can’t kill the bacteria. Many bacteria develop resistance by restricting entry.
Porin Changes: P. aeruginosa can alter its porins (channel proteins), effectively closing the “doors” to tetracyclines.
Natural Barriers: Some bacteria, like Streptococci, have a natural permeability barrier that makes them inherently resistant to aminoglycosides.
5. Alteration of Metabolic Pathways
If an antibiotic blocks a specific chemical reaction the bacteria need to survive, some bacteria simply find a “detour.”
Example: Sulfonamides work by preventing bacteria from making folic acid from PABA. However, some resistant bacteria have evolved to bypass this requirement entirely by utilizing preformed folic acid from their environment, much like human cells do.
6. Efflux Pumps: The Cellular “Ejector Seat”
Efflux pumps are specialized proteins located in the bacterial cell membrane that actively pump antibiotics out of the cell before they can reach toxic levels.
Major Families: These include the ABC (ATP-binding cassette) pumps, MATE (multidrug resistance and toxic compound extrusion) pumps, and the MFSE (major facilitator superfamily efflux) pumps.
These pumps are often responsible for resistance to multiple classes of drugs, including quinolones and aminoglycosides.
How Resistance Spreads: Bacterial Genetics
Bacteria do not just inherit resistance from their “parents”; they can also share resistance genes with their “neighbors” like a digital file transfer.
Vertical Gene Transfer (VGT)
This is generational inheritance. When a resistant bacterium divides, it passes its resistant genes down to its daughter cells. Through rapid reproduction, a single resistant cell can create a colony of millions in a matter of hours.
Horizontal Gene Transfer (HGT)
This is the most alarming aspect of antibiotic resistance, as it allows resistance to jump between different species of bacteria.
Transformation: Bacteria pick up fragments of DNA containing resistance genes from their surrounding environment (often left behind by dead bacteria).
Transduction: Resistance genes are carried from one bacterium to another by a virus known as a bacteriophage.
Conjugation (Plasmid Transfer): Often called “bacterial sex,” this involves the direct transfer of circular DNA molecules called plasmids between bacteria. These plasmids often carry “resistance cassettes” that contain genes for resistance to multiple different antibiotics at once.
The Global Impact and the Need for Action
The statistics are sobering. Hundreds of thousands of people die every year globally from infections that were once easily treatable.
Note: If left unchecked, it is estimated that antimicrobial resistance could cause 10 million deaths annually by 2050, surpassing cancer as a leading cause of death.
The Root of the Problem
The rapid increase in resistance is primarily driven by:
Overuse: Prescribing antibiotics for viral infections (like the common cold or flu) where they have zero effect.
Misuse: Patients failing to complete their full course of antibiotics, which allows the strongest bacteria to survive and develop resistance.
Agricultural Use: The widespread use of antibiotics in livestock to promote growth rather than treat disease.
The Path Forward
To combat this “silent pandemic,” the global community must intensify efforts in two key areas:
New Drug Development: We must incentivize the discovery of new classes of antibiotics and alternative therapies, such as phage therapy or monoclonal antibodies.
Stewardship: Implementing strict “antibiotic stewardship” programs in hospitals to ensure the right drug is used at the right dose for the right duration.
Summary Table: Mechanisms of Resistance
| Mechanism | Description | Example |
| Enzyme Production | Destroys the drug molecule | beta-lactamase vs. Penicillin |
| Altered Enzymes | Enzyme changes shape to avoid drug binding | Modified Dihydrofolate reductase |
| Target Modification | Changes the drug’s “landing site” | PBP alteration in S. pneumoniae |
| Permeability Change | Closes entry ports in the cell wall | Porin changes in P. aeruginosa |
| Metabolic Bypass | Finds a new way to make essential nutrients | Utilizing preformed folic acid |
| Efflux Pumps | Actively pumps the drug out of the cell | ABC and MATE pumps |
Conclusion
Bacterial resistance is a natural evolutionary process, but human activity has accelerated it to a dangerous pace. By understanding the complex mechanisms—from enzyme production to efflux pumps—and recognizing how genetic material is swapped through transformation and transduction, we can better appreciate the magnitude of this challenge. The future of modern medicine depends on our ability to outsmart these microscopic adversaries through innovation, education, and responsible drug use.