Abstract
The tumor suppressor protein p53 is a master regulator of cellular stress responses, orchestrating DNA repair, cell cycle arrest, senescence, and apoptosis. Owing to its pivotal role in maintaining genomic integrity, TP53 is the most frequently mutated gene in human cancers. This review provides a detailed, publication-ready overview of p53 structure, regulatory mechanisms, activation pathways, and mechanisms of inactivation in malignancies, with emphasis on translational relevance and therapeutic targeting.
1. Introduction
The p53 protein, widely known as the “guardian of the genome,” is a sequence-specific transcription factor that preserves genomic stability. It responds to diverse cellular stresses—including DNA damage, oncogene activation, and hypoxia—by regulating gene expression programs that determine cell fate. Loss or dysfunction of p53 is a defining feature of tumorigenesis, underscoring its importance in cancer biology.
2. Structural Organization of p53

p53 functions as a tetramer and consists of five key functional domains:
Transactivation Domain (TAD): Initiates transcription of downstream target genes.
Proline-Rich Domain (PRD): Facilitates apoptotic signaling and protein interactions.
DNA-Binding Domain (DBD): Responsible for sequence-specific DNA binding; the most frequent site of cancer-associated mutations.
Oligomerization Domain (OD): Enables tetramer formation, which is essential for transcriptional activity.
C-terminal Regulatory Domain: Modulates DNA binding and undergoes post-translational modifications.
Highly conserved regions (Boxes I–V) across species highlight the evolutionary significance of p53 function.
3. Regulation of p53 Stability and Activity
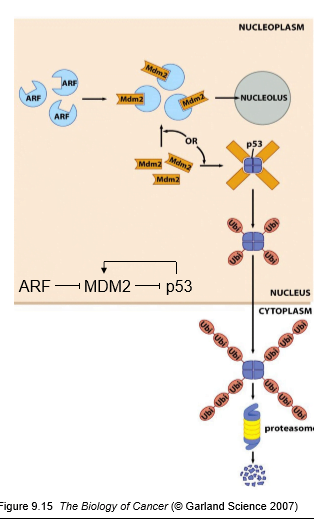
Figure 2. MDM2-Mediated Negative Feedback Loop
p53 is a short-lived protein with a half-life of approximately 5–20 minutes under basal conditions. Its intracellular levels are primarily controlled through ubiquitin-mediated degradation rather than synthesis.
Key Regulatory Mechanisms
MDM2 as an E3 ubiquitin ligase: Targets p53 for proteasomal degradation.
Nuclear export: MDM2 promotes cytoplasmic localization of p53.
Negative feedback loop: p53 transcriptionally activates MDM2, maintaining homeostasis.
ARF-mediated stabilization: ARF inhibits MDM2, preventing p53 degradation and enhancing its activity.
This tightly controlled regulatory network ensures that p53 is activated only under stress conditions, preventing unnecessary cell cycle arrest or apoptosis.
4. Pathways Leading to p53 Activation
Figure 3. Major p53 Activation Pathways
DNA Damage → ATM/Chk2 → p53 phosphorylation → stabilization
UV Stress → ATR → p53 activation
Oncogenes (Ras/Myc) → ARF → MDM2 inhibition → p53 activation
p53 is activated through multiple signaling pathways in response to cellular stress:
4.1 DNA Damage Response
Ionizing radiation induces DNA double-strand breaks, activating ATM and Chk2 kinases. These kinases phosphorylate p53, disrupting its interaction with MDM2 and stabilizing the protein.
4.2 UV Radiation and Replication Stress
UV radiation activates ATR kinase, which similarly phosphorylates p53 and enhances its transcriptional activity.
4.3 Oncogene-Induced Stress
Oncogenes such as Ras and Myc activate ARF, which inhibits MDM2 and stabilizes p53. This serves as a critical safeguard against uncontrolled proliferation.
5. Functional Outcomes of p53 Activation
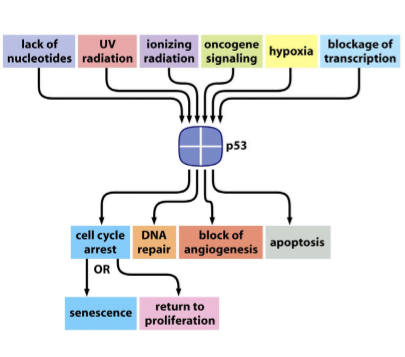
Figure 4. Cellular Outcomes of p53 Activation
p53 Activation
|
---------------------------------------
| | |
Cell Cycle DNA Repair Apoptosis
Arrest (G1) (Cell Death)
Upon activation, p53 determines cellular fate based on the severity of damage and cellular context:
5.1 Cell Cycle Arrest
p53 induces expression of p21, leading to G1 phase arrest. This pause allows cells to repair damaged DNA before replication.
5.2 DNA Repair
p53 activates genes involved in DNA repair pathways, promoting genomic stability.
5.3 Apoptosis
In cases of severe or irreparable damage, p53 induces apoptosis by activating pro-apoptotic genes such as BAX and PUMA.
The balance between these outcomes is influenced by cell type, stress intensity, and availability of survival signals.
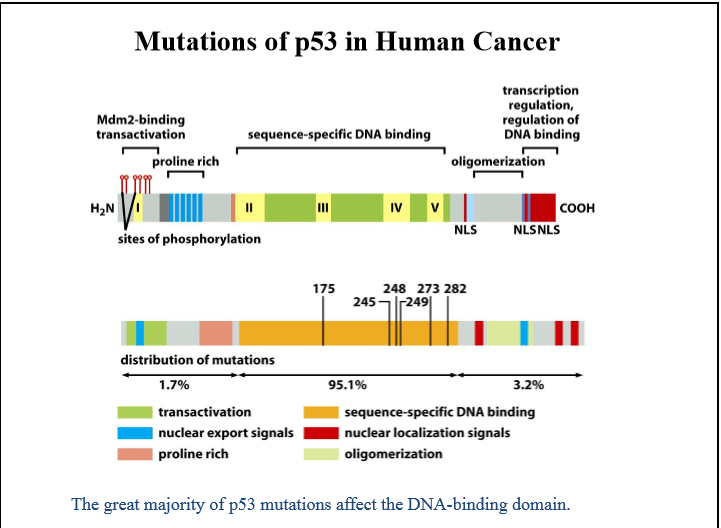
6. Mutations of p53 in Human Cancer
TP53 is mutated in approximately 50% of human cancers, making it the most commonly altered tumor suppressor gene.
Common Features of p53 Mutations
Predominantly missense mutations
Frequently occur in the DNA-binding domain
Result in loss of transcriptional activity
Consequences of Mutation
Loss of tumor suppressor function
Increased genomic instability
Resistance to apoptosis
Potential gain-of-function effects promoting tumor progression
7. Mechanisms of p53 Inactivation in Cancer
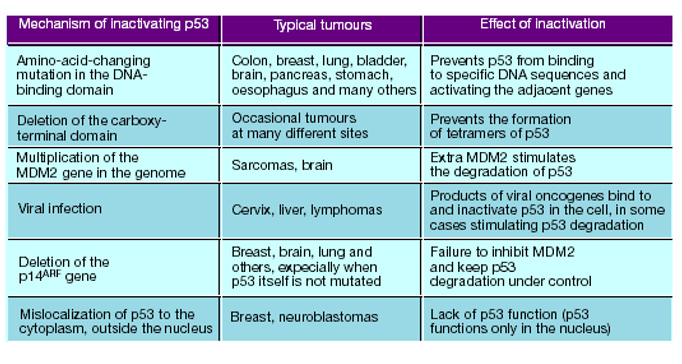
Figure 5. Mechanisms of p53 Inactivation
TP53 mutation
MDM2 overexpression
Loss of ARF function
Viral proteins (e.g., HPV E6)
Beyond direct mutation, p53 can be inactivated through several mechanisms:
MDM2 overexpression: Leads to excessive degradation of p53.
ARF loss: Removes inhibition of MDM2, reducing p53 stability.
Viral oncogenes: Proteins such as HPV E6 target p53 for degradation.
These mechanisms collectively contribute to tumor development and progression.
8. Therapeutic Targeting of the p53 Pathway
Given its central role in cancer, the p53 pathway is a major focus of therapeutic development:
MDM2 inhibitors (e.g., Nutlin-3): Restore p53 function by preventing its degradation.
Mutant p53 reactivators: Aim to restore wild-type conformation and function.
Gene therapy approaches: Deliver functional TP53 to tumor cells.
These strategies hold promise for improving outcomes in cancers with p53 pathway dysfunction.
9. Discussion
The p53 signaling network represents one of the most complex and tightly regulated systems in cell biology. Its ability to integrate multiple stress signals and determine cell fate underscores its importance in preventing malignant transformation. However, this complexity also presents challenges for therapeutic targeting, as context-dependent effects must be carefully considered.
10. Conclusion
p53 is a cornerstone of tumor suppression, regulating critical processes such as cell cycle arrest, DNA repair, and apoptosis. Its structural domains, regulatory mechanisms, and activation pathways collectively ensure genomic stability. Disruption of p53 function is a hallmark of cancer, making it a key target for therapeutic intervention.
11. Future Perspectives
Future research should focus on understanding mutant p53 gain-of-function activities, improving targeted therapies, and integrating p53-based strategies into precision oncology.