RNA interference (RNAi) is a biological process. It is defined as the post-transcriptional gene silencing process mediated by short double-stranded RNA molecules called siRNA (small interfering RNAs) and micro RNA(miRNA) by neutralizing targeted mRNA molecules.
Andrew Fire and Craig C. Mello shared the 2006 Nobel Prize in Medicine for their work on RNAi in the nematode worm C. elegans.
History of RNAi
RNAi was known by other names, including co-suppression, post-transcriptional gene silencing (PTGS), and quelling, historically. RNA interference (RNAi) was discovered unexpectedly during attempts to experimentally manipulate the expression of specific genes. Researchers tried to inhibit the expression of a gene in C. elegans by microinjecting a single-stranded, complementary RNA that would hybridize to the encoded mRNA and prevent its translation, a method called antisense inhibition. But in control experiments, perfectly base-paired double-stranded RNA a few hundred base pairs long was much more effective at inhibiting expression of the gene than the antisense strand alone. Thus, RNAi was initially discovered and characterized in the nematode worm, C. elegans. The double-stranded RNA (dsRNA) was found to be 10-times more effective in silencing target gene expression than antisense or sense RNA alone from the studies. Similar inhibition of gene expression by an introduced double-stranded RNA soon was later observed in plants.
Components of the RNAi pathway
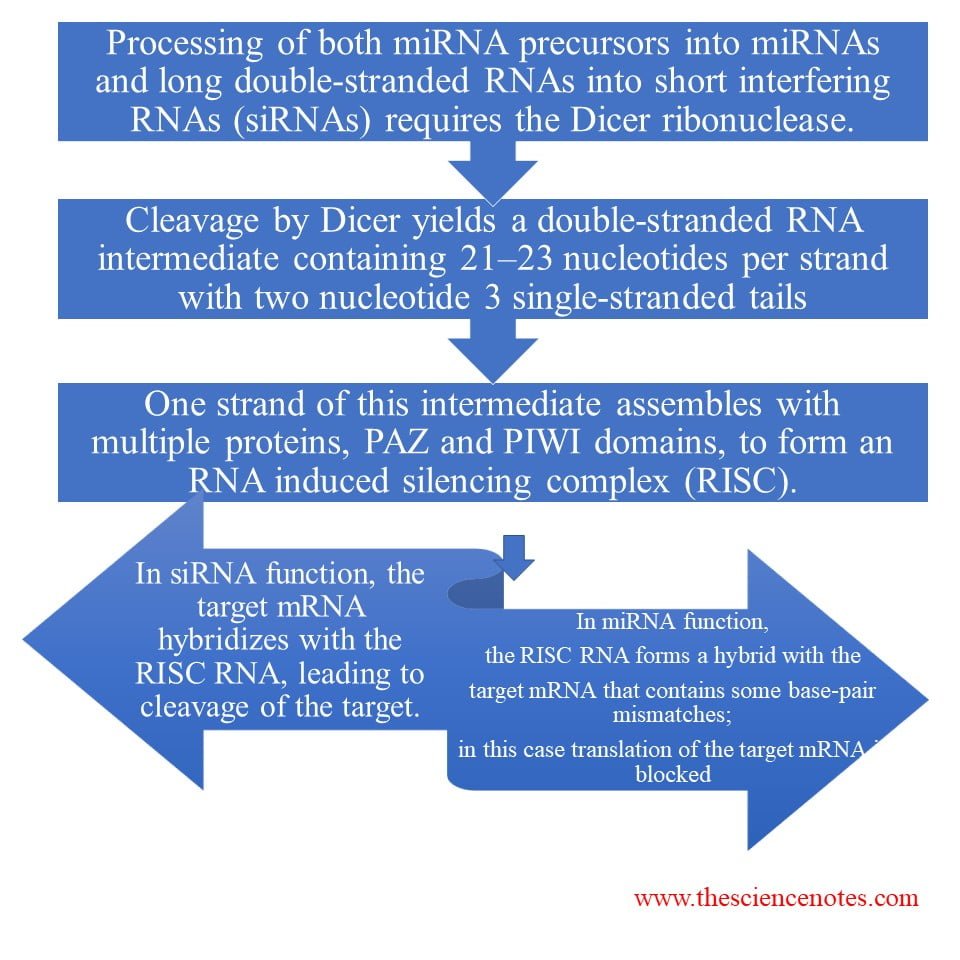
1.Dicer family proteins
Dicer family proteins contain an N-terminal helicase domain, a C-terminal segment containing dual RNase III domains, and one or more dsRNA-binding motifs. Family members also contain a PAZ domain. Dicer forms short duplexes about 20 to 25 nucleotides long. One strand of the processed miRNA is transferred to the target mRNA (or to a viral or transposon RNA), leading to inhibition of translation or degradation of the RNA.
2.Argonaute (RISC complex):
Argonaute family members are highly basic, ~100 kD proteins that contain PAZ and PIWI domains.
3.Small non-coding RNA; microRNAs (miRNAs) and small interfering RNAs (siRNAs)
Small non-coding RNA; microRNAs (miRNAs) and small interfering RNAs (siRNAs; Both varieties are pieces of RNA – approximately 22 nucleotides long – but they differ in their role, specificity and how they are synthesized.
MicroRNA (miRNA)
miRNAs are a more general suppressive tool, special class of single-stranded RNA precursors and characterized by their distinctive hairpin shape found in all eukaryotes. They are noncoding RNAs, about 22 nucleotides long, complementary in sequence to particular regions of mRNAs. They regulate mRNA function by cleaving the mRNA or suppressing its translation. Up to 1% of the human genome may encode miRNAs, and miRNAs may target up to one-third of human mRNAs.
Small interfering RNA (siRNA)
siRNAs are highly specific and usually synthesized to reduce the translation of specific messenger RNAs (mRNAs) for inducing short-term silencing of protein coding genes. This is done to reduce the synthesis of particular proteins. They are formed from double-stranded RNA transcribed and then cut to size in the nucleus before releasing into the cytoplasm. Small interfering RNAs (siRNAs) are knoen to be the most promising type of RNA-based therapeutic oligonucleotide drug, since their mechanism of action is catalytic and each siRNA molecule can inactivate several target RNA molecules in a sequence-specific manner.
How RNAi Works?
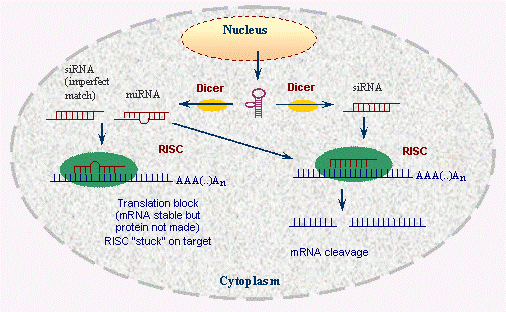
Regardless of whether siRNAs or miRNAs are involved, RNA interference works in the same process. These small RNA molecules connect to and activate protein complexes, most notably the RNA-induced silencing complex (RISC). After binding to their target mRNAs and both physically prevent ribosomes from continuing to synthesize the associated protein and mark that mRNA for destruction or repression. They are also known to direct the formation of heterochromatin for further target RNA transcription.

Role of RNAi
Since, RNA interference is a natural process with a role in the regulation of protein synthesis and in immunity, they can have numerous role in modern molecular technology.Some of them are highlighted below;
- Genome Wide Regulation; RNAi plays a role in regulating development and genome maintenance.30% of human genome regulated. It’s also a potent tool for the exploration and manipulation of gene expression. Its targets include RNA from viruses and transposons.
- Defense Mechanism; RNA interference also serves a role in protecting cells from viruses, by attacking their mRNAs and sometimes even their RNA-based genomes before the cell’s synthesis machinery can manufacture viral proteins.
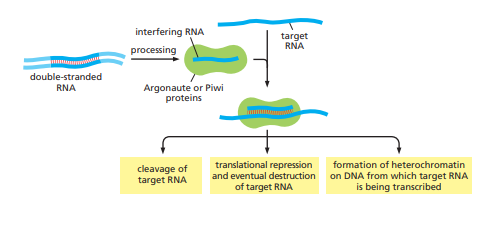
[Source; Alberts, Bruce, Alexander Johnson, Julian Lewis, Martin Raff, Keith Roberts, and Peter Walter. Molecular Biology of the Cell]
- Research: These RNAs then inhibit protein synthesis and the genetic code.A protein’s function is still unknown, iRNA biology helps scientists to enable “knockdown” experiments, in which synthesis of a protein is reduced but not eliminated, as it would be if its gene were simply cut out or turned off.
- Medicine: Since the mRNAs is targeted rather than proteins, RNA interference stands to work around the cross-reactivity and resistance that can be helpful for complicated treatment for many conditions making more specific targets while reducing side effects. RNA interference is a fertile area of inquiry in many sub-fields of medicine. It may become an important weapon against increasingly antibiotic-resistant bacterial infections, inhibiting bacteria in an entirely different way than antibiotics do. Therapeutic uses of RNAi in hematology(blood),loss of gene function, oncology(cancer),Stem cell biology and infectious diseases is known so far.
Learn more:
References
- Lehninger, Albert L., Cox, Michael M. Nelson, David L. Lehninger Principles Of Biochemistry. New York : W.H. Freeman, 2008 pp 1215-1218
- Alberts, Bruce, Alexander Johnson, Julian Lewis, Martin Raff, Keith Roberts, and Peter Walter. Molecular Biology of the Cell. New York: Garland Science, 2002.pp 430-135
- Lodish, Harvey F. Molecular Cell Biology. New York: W.H. Freeman and Co, 5th edition.pp 518-525
- Dana, H., Chalbatani, G. M., Mahmoodzadeh, H., Karimloo, R., Rezaiean, O., Moradzadeh, A., Mehmandoost, N., Moazzen, F., Mazraeh, A., Marmari, V., Ebrahimi, M., Rashno, M. M., Abadi, S. J., & Gharagouzlo, E. (2017). Molecular Mechanisms and Biological Functions of siRNA. International journal of biomedical science : IJBS, 13(2), 48–57.
- Agrawal, N., Dasaradhi, P. V., Mohmmed, A., Malhotra, P., Bhatnagar, R. K., & Mukherjee, S. K. (2003). RNA interference: biology, mechanism, and applications. Microbiology and molecular biology reviews : MMBR, 67(4), 657–685. https://doi.org/10.1128/mmbr.67.4.657-685.2003
- Obbard, D. J., Gordon, K. H., Buck, A. H., & Jiggins, F. M. (2009). The evolution of RNAi as a defence against viruses and transposable elements. Philosophical transactions of the Royal Society of London. Series B, Biological sciences, 364(1513), 99–115. https://doi.org/10.1098/rstb.2008.0168
- Chernikov Ivan V., Vlassov Valentin V., Chernolovskaya Elena L.(2019).Current Development of siRNA Bioconjugates: From Research to the Clinic. Frontiers in Pharmacology .Vol.10. pp 444 .DOI=10.3389/fphar.2019.00444
- https://www.iomcworld.org/medical-journals/rna-interference-52137.html
- https://www.ncbi.nlm.nih.gov/probe/docs/techrnai/
- https://www.sciencedirect.com/topics/neuroscience/small-interfering-rna
- https://www.umassmed.edu/rti/biology/how-rnai-works/
- https://www.thermofisher.com/blog/ask-a-scientist/what-is-rnai/