Abstract
- TCR or T-cell receptors is the antigen receptor present on T-cells. It is structurally similar to BCR or B-cell receptors.
- T and B cells are lymphocytic cells which uses various chains to recognize antigens. TCR can bind to or recognize antigens bound to MHC or major histocompatibility complex on the surface of APC or antigen presenting cells.
- It is first processed by APC and then presented. There are various T-cell accessory proteins helping TCR. There are also various adaptor proteins with specific domains. When TCR is activated alteration of gene transcription occurs.
- TCR diversity is very important in recognizing the wise range of antigens present. TCR signaling errors can lead to development various immunological problems also. The mutations in the adaptor proteins and alterations in TCR genes can cause this. But TCR based immunotherapy is also developing. Various technologies using adoptive cell therapy (ACT), Chimeric antigen receptor (CARs) and soluble T-cell receptors are being used.
What are T cell receptors?
- TCR or T-cell receptors are basically groups of polypeptide chains found on the surface of T lymphocytes. Lymphocytes are a type of white blood cell consisting of natural killer cells, T lymphocytes and B lymphocytes.
- T lymphocytes are a specific type of lymphocytes which develops in thymus gland whereas B lymphocytes are bone marrow or bursa derived cells and are part of the immune response system of the body (Richard A. Goldsby, Thomas J. Kindt, Janis Kuby, 2002).
Classes of T-cell receptors
- One is α: β T-cell having α and β chains and the
- other is γ: δ T-cell having γ and δ chains.
The first type is most common in humans. Almost in 95% of T-cells TCR have α and β chain. The genes are termed as TRA, TRB. The rest 5% consists of γ and δ chains encoded by TRG and TRD.
In the year 1984 TCR was discovered. Eventually the entity and structure of TCR was discovered which finally led to the important studies in the fields of CAR-T, cancer immunotherapy and checkpoint inhibition (Filion, 1984; Yanagi et al., 1984). Chimeric antigen receptors (CAR) are fusion proteins engineered from antigen recognition, signaling, and costimulatory domains that can be used to reprogram T cells to specifically target tumor cells expressing specific antigens (Petty et al., 2020). T cells can respond to antigenic ligands derived from pathogens but can remain inert to similar ligands from host tissues. The behavior of the signaling receptors are responsible for this (Adam H. Courtney, Wan-Lin Lo, 2018). The specificity of T-cell is determined by TCR. Each T-cell has TCR with unique specificity. Short fragments of peptide bound to MHC class I or MHC class II molecules also called as pMHC are surveyed by TCR (Adam H. Courtney, Wan-Lin Lo, 2018). Each T cell possesses a TCR of unique specificity but TCR may interact with different types of MHC ligand (Sewell, 2012). There are mainly three types of T cells – helper T cells, regulatory T cells and cytotoxic T cells. T helper cells or Th cells are also called as CD4+ cells. They help activity of cells involved with immunity of body. Th cells which are mature expresses CD4 on their surface and so called as CD4+ cells. In HIV infection active CD4+ cells decrease which leads to AIDS. T-regulatory cells also called as suppressor cells are generally immunosuppressive which controls overexpression of effector T-cells (Bettelli et al., 2006). Cytotoxic T-cells are the T lymphocytes capable of killing cancer cells. It is accompanied by CD8 glycoprotein which is also required for its binding to MHC and so are called as CD8+ T cells. It has the ability to form cytokines also. TCR stimulation can lead to T-cell differentiation also. There is naïve T-cell from which activated T-cell, effector T-cell and memory T-cell forms.
Structure and T cell receptor signalling
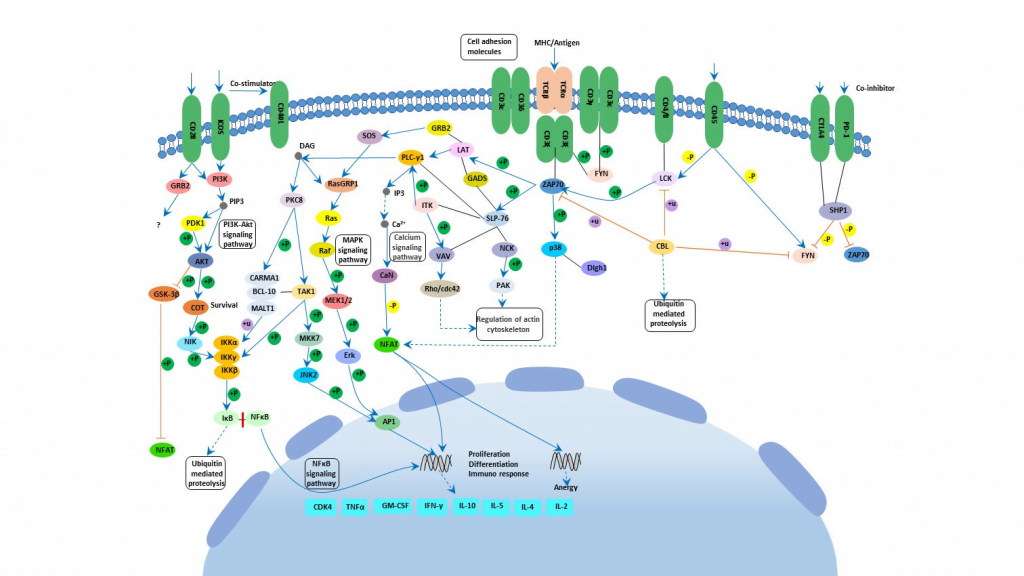
As discussed there are two types of TCR based on their chains. Each chain has two distinct regions named variable region and constant region. Variable region has 3 hypervariable region called complementary determining regions or CDR. There is also another region on β which is not considered CDR as it does not interact with antigen. CDR1 interacts with the N and C terminals of the antigenic peptides. CDR3 is mainly responsible for attaching to peptides. Like Fab of immunoglobulins T-cell receptors acts as the antigen receptor on T cells. The MHC-TCR-CD3 interaction in T cells are like Ag-Ig-CD79 interaction in B cells. T cells express invariant chains comprising of the CD3 complex. Cluster of differentiation 3 or CD3 helps in activating cytotoxic T cells and helper T cells. There are 4 chains – CD3δ, CD3γ, CD3ε, CD3ζ (Kenneth Murphy, Paul Travers, 2008). Cytoplasmic domain of CD3 and ζ chains are required for signaling. The TCR complex is basically formed by TCR, CD3 and the ζ chain. They contain a tyrosine embedded conserved amino acid motif called immunoreceptor tyrosine-based activation motif or ITAM. These complex structure was found using CryoEM (Dong et al., 2019). When ITAM is phosphorylated it can recruit other signaling proteins. There are some proximal TCR signaling molecule Zap70, Lck, LAT etc. When pMHC bind to TCR, Lck is recruited to TCR complex. Lck then phosphorylates ITAM (Adam H. Courtney, Wan-Lin Lo, 2018). ITAM motifs have tyrosines which creates binding site for SH2 domain of Zap70 kinase after phosphorylation. Activation of Zap70 occurs and is phosphorylated by Lck. Zap again in turn phosphorylates LAT and SLP-76 which are two adapter proteins. As a result of this, activation of phospholipase C gamma or PLC-γ occurs. This PLC-γ can cleave PIP2 or phosphatidylinositol bisphosphate to form DAG or diacylglycerol and IP3 or inositol trisphosphate. DAG activates protein kinase C or PKC leading to activation of transcription factor NFκB. IP3 on the other hand activates a phosphatase calcineurin which activates NFAT or nuclear factor of activated T cells. DAG also plays role in activating the MAP kinase cascade which finally activates AP-1 which is another transcription factor. All this transcription factors induces gene transcription leading to cell proliferation (Kenneth Murphy, Paul Travers, 2008)(Adam H. Courtney, Wan-Lin Lo, 2018). There are co receptors which enhances the signal from T cell. They bind to the MHC molecules. Co-receptor CD4 which is specific for MHC class II helps helper and regulatory T cells whereas co-receptor CD8 is specific for MHC class I and it helps cytotoxic T cells. The co-receptors recruit essential molecules like Lck.
Generation of TCR diversity is very important for recognizing different types of antigens. For effective control of viral infection also it is very important (Chen et al., 2012; Messaoudi et al., 2002). Variable region of TCR α and TCR δ have number of variable (V) genes whereas TCR β and TCR γ have an additional diversity (D) genes along with V genes (Miles et al., 2011)(Rosati et al., 2017). During VDJ recombination alleles of each gene recombines with other and forms a functional variable region. Strong combinatorial and junctional diversity develops resulting in formation of highly variable TCR receptor which can identify plethora of antigens. Pairing of α and β gives additional diversity (Turner et al., 2006)(Rosati et al., 2017). Within the variable domain variable (V) region have CDR1 and CDR2. CDR3 has D and J along with some of V region. The most variable is CDR3 (William E. Paul, 2008). Each recombined TCR have unique specificity towards antigens. The antigen binding site is also responsible and it is based on the α:β T-cell or γ:δ T-cell(Kenneth Murphy, Paul Travers, 2008). TCR α, γ chain diversity derives from VJ recombination and TCR β, δ involves VDJ recombination. The total diversity of immunoglobulin is around 5*10^13 and α:β T-cell receptor is around 10^18 (Peter Parham, 2009).
TCR signaling errors
There can be mutations in the genes involved in TCR signaling leading to various immune pathologies and immune dysregulation. Mutation in Zap70, Lck, ITK leads to various clinical difficulties (Luigi D. Notarangelo, 2014). It was seen that CD8 deficiency can lead to mutation in Zap70 gene (Arpaia et al., 1994; Chan et al., 1994; Elder et al., 1994). Lack of circulating CD8+ T cells can lead to CD8 deficiency. CD4+ cells from Zap70 mutated patients shows reduced level of CTLA4, IL-10, TGFβ. Sensitivity to Fas mediated apoptosis also decreases (Roifman et al., 2010). Patient with CD8 deficiency shows unique CD4 T cells which leads to infiltrative cutaneous lesions and accumulation of eosinophil in perivascular area of skin (Katamura et al., 1999). Also it was seen that missense mutation in Zap70 can lead to erythroderma, eosinophilia and elevated serum IgE levels (Turul et al., 2008). Zak related SCID has almost 12 mutations. Lck deficiency can lead to respiratory infections and formation of skin lesion (Luigi D. Notarangelo, 2014). Missense mutation in Lck gene gives rise to defective protein which lacks kinase activity. Also in Lck deficient patients CD4+ T cells and regulatory T cells were found to be less (Hauck et al., 2012). ITK deficiency can also lead to clinical problem as it is an important part of TCR activation and regulation. It promotes PLC-γ activation. Deficiency of ITK leads to immune pathology. Various lung disease can occur due to this. Deficiency of natural killer cells and improper CD3 stimulation is seen in patients having ITK deficiency. Apart from this there can be immune dysregulation due to reduced TCR signaling, abnormal regular T-cell development, defect in T-cell survival. In mice it has been seen that missense mutation in SLP-76 causes impaired intrathymic deletion of self-reactive CD4 cells (Luigi D. Notarangelo, 2014).
Therapies mediated by T-cell
T-cell based immunotherapy is one important way by which therapies against cancer can be made. Various research has been carried out. It is emerging as a potential breakthrough in cancer therapeutics. Among the various types of T-cell therapies, one is adoptive cell therapy. It is based on transfer of cells (Tran et al., 2008). Isolation of T-cells from patient suffering from cancer is done followed by in-vitro culture of the cells. Finally, it is reintroduced into the same patient. This is also called as autologous cancer immunotherapy. In allogenic therapy different donor and receiver is there (Marcus & Eshhar, 2011).
Sometimes the T-cells are genetically engineered for the production of artificial T-cell receptors (Jensen et al., 2019). This are called as Chimeric antigen receptor T cells or CAR-T cells. The basis of CAR-T is modification of T-cells which recognizes cancer cells so that it can more effectively identify and destroy the cancer cells. After harvesting T-cells from patients it is genetically altered and infused into cancer patients. It can be both autologous and allogenic. CAR-T cells act as living drug against cancerous cells (Sadelain et al., 2013). The cells perform based on their cytotoxicity. They increase the production of compounds which may affect cell proliferation like cytokines, interleukins and growth factors (Tang et al., 2015). Cytokine interleukins and anti CD3 antibodies are used to expand T-cells (Makita et al., 2017). Gamma-retroviruses and lentiviruses with partially deleted U3 region are used to integrate the CAR genes into specific site of genome (Jensen et al., 2019; Jin et al., 2016). Researches are going on to develop CARs which may work in Hodgkin’s lymphoma, acute myeloid leukemia etc. (Schultz & Mackall, 2019). The inability to control cytokine release and tumor lysis, the absence of an off switch are some of the challenges of the CAR-T cell therapies. Use of soluble T-cell receptor therapy is another alternative. A full length TCR have a transmembrane region which is not present in soluble TCR. It has antigen recognition parts. It can be used for antigen specific labeling and elimination of target cells of human beings (Walseng et al., 2015).
Future perspectives
It has been seen that T-cell mediated immune response can control formation of tumor. Various antigens on tumors have been identified also (Coulie et al., 2001; de Aquino et al., 2015; Van Der Bruggen et al., 2002). But there are various challenges like self-nature of the tumor antigens as a result of which TCR have low affinity. To have active T-cell activated novel peptides can play a role (Seidel et al., 2012). Suppressor factors has to be blocked and there should be availability of CD4+ T-cell. Tumors can alter surface expression preventing T-cells from binding. Abnormal TCR-CD3 complex can form due to tumor growth and TCR signal gets hampered. Sometimes TCR-MHC interaction also gets inhibited due to tumor formation (de Aquino et al., 2015). So future efforts should focus on these issues for the production of a successful TCR based immunotherapy which can help in curing the immune diseases as well as help cancer therapy.
References
- Adam H. Courtney, Wan-Lin Lo, and A. W. (2018). HHS Public Access. Trends Biochem Sci., 43(2), 108–123. https://doi.org/10.1016/j.tibs.2017.11.008
- Arpaia, E., Shahar, M., Dadi, H., Cohen, A., & Rolfman, C. M. (1994). Defective T cell receptor signaling and CD8+ thymic selection in humans lacking Zap-70 kinase. Cell, 76(5), 947–958. https://doi.org/10.1016/0092-8674(94)90368-9
- Bettelli, E., Carrier, Y., Gao, W., Korn, T., Strom, T. B., Oukka, M., Weiner, H. L., & Kuchroo, V. K. (2006). Reciprocal developmental pathways for the generation of pathogenic effector TH17 and regulatory T cells. Nature, 441(7090), 235–238. https://doi.org/10.1038/nature04753
- Chan, A. C., Kadlecek, T. A., Elder, M. E., Filipovich, A. H., Kuo, W. L., Iwashima, M., Parslow, T. G., & Weiss, A. (1994). ZAP-70 deficiency in an autosomal recessive form of severe combined immunodeficiency. Science, 264(5165), 1599 LP – 1601. https://doi.org/10.1126/science.8202713
- Chen, H., Ndhlovu, Z. M., Liu, D., Porter, L. C., Fang, J. W., Darko, S., Brockman, M. A., Miura, T., Brumme, Z. L., Schneidewind, A., Piechocka-Trocha, A., Cesa, K. T., Sela, J., Cung, T. D., Toth, I., Pereyra, F., Yu, X. G., Douek, D. C., Kaufmann, D. E., … Walker, B. D. (2012). TCR clonotypes modulate the protective effect of HLA class I molecules in HIV-1 infection. Nature Immunology, 13(7), 691–700. https://doi.org/10.1038/ni.2342
- Coulie, P. G., Hanagiri, T., & Takenoyama, M. (2001). From tumor antigens to immunotherapy. International Journal of Clinical Oncology, 6(4), 163–170. https://doi.org/10.1007/PL00012101
- de Aquino, M. T. P., Malhotra, A., Mishra, M. K., & Shanker, A. (2015). Challenges and future perspectives of T cell immunotherapy in cancer. Immunology Letters, 166(2), 117–133. https://doi.org/10.1016/j.imlet.2015.05.018
- Dong, D., Zheng, L., Lin, J., Zhang, B., Zhu, Y., Li, N., Xie, S., Wang, Y., Gao, N., & Huang, Z. (2019). Structural basis of assembly of the human T cell receptor–CD3 complex. Nature, 573(7775), 546–552. https://doi.org/10.1038/s41586-019-1537-0
- Elder, M. E., Lin, D., Clever, J., Chan, A. C., Hope, T. J., Weiss, A., & Parslow, T. G. (1994). Human severe combined immunodeficiency due to a defect in ZAP-70, a T cell tyrosine kinase. Science, 264(5165), 1596 LP – 1599. https://doi.org/10.1126/science.8202712
- Filion, L. (1984). {\copyright} 1984 Nature Publishing Group. Group, 311, 276–279.
- Hauck, F., Randriamampita, C., Martin, E., Gerart, S., Lambert, N., Lim, A., Soulier, J., Maciorowski, Z., Touzot, F., Moshous, D., Quartier, P., Heritier, S., Blanche, S., Rieux-Laucat, F., Brousse, N., Callebaut, I., Veillette, A., Hivroz, C., Fischer, A., … Picard, C. (2012). Primary T-cell immunodeficiency with immunodysregulation caused by autosomal recessive LCK deficiency. Journal of Allergy and Clinical Immunology, 130(5), 1144-1152.e11. https://doi.org/10.1016/j.jaci.2012.07.029
- Jensen, T. I., Axelgaard, E., & Bak, R. O. (2019). Therapeutic gene editing in haematological disorders with CRISPR/Cas9. British Journal of Haematology, 185(5), 821–835. https://doi.org/10.1111/bjh.15851
- Jin, C., Fotaki, G., Ramachandran, M., Nilsson, B., Essand, M., & Yu, D. (2016). Safe engineering of CAR T cells for adoptive cell therapy of cancer using long-term episomal gene transfer. EMBO Molecular Medicine, 8(7), 702–711. https://doi.org/10.15252/emmm.201505869
- Katamura, K., Tai, G., Tachibana, T., Yamabe, H., Ohmori, K., Mayumi, M., Matsuda, S., Koyasu, S., & Furusho, K. (1999). Existence of activated and memory CD4+ T cells in peripheral blood and their skin infiltration in CD8 deficiency. Clinical and Experimental Immunology, 115(1), 124–130. https://doi.org/10.1046/j.1365-2249.1999.00759.x
- Kenneth Murphy, Paul Travers, and M. W. (2008). The Seventh Edition of the Janeway’s Immunobiology. Garland Science, 1, 865. https://doi.org/10.1111/j.1365-3083.2008.02123.x
- Luigi D. Notarangelo. (2014). NIH Public Access. Curr Opin Immunol, 0, 97–101. https://doi.org/10.1016/j.coi.2014.10.003
- Makita, S., Yoshimura, K., & Tobinai, K. (2017). Clinical development of anti-CD19 chimeric antigen receptor T-cell therapy for B-cell non-Hodgkin lymphoma. Cancer Science, 108(6), 1109–1118. https://doi.org/10.1111/cas.13239
- Marcus, A., & Eshhar, Z. (2011). Allogeneic adoptive cell transfer therapy as a potent universal treatment for cancer. In Oncotarget (Vol. 2, Issue 7, pp. 525–526). https://doi.org/10.18632/oncotarget.300
- Messaoudi, I., Patiño, J. A. G., Dyall, R., LeMaoult, J., & Nikolich-Žugich, J. (2002). Direct Link Between <em>mhc</em> Polymorphism, T Cell Avidity, and Diversity in Immune Defense. Science, 298(5599), 1797 LP – 1800. https://doi.org/10.1126/science.1076064
- Miles, J. J., Douek, D. C., & Price, D. A. (2011). Bias in the αΒ T-cell repertoire: Implications for disease pathogenesis and vaccination. Immunology and Cell Biology, 89(3), 375–387. https://doi.org/10.1038/icb.2010.139
- Peter Parham. (2009). The Immune System. In Garland Science. YJBM. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4345546/
- Petty, A. J., Heyman, B., & Yang, Y. (2020). Chimeric Antigen Receptor Cell Therapy : Overcoming Obstacles to Battle Cancer. 40, 1–18. https://doi.org/10.3390/cancers12040842
- Richard A. Goldsby, Thomas J. Kindt, Janis Kuby, B. A. O. (2002). Kuby Immunology.pdf.
- Roifman, C. M., Dadi, H., Somech, R., Nahum, A., & Sharfe, N. (2010). Characterization of ζ-associated protein, 70 kd (ZAP70)–deficient human lymphocytes. Journal of Allergy and Clinical Immunology, 126(6), 1226-1233.e1. https://doi.org/10.1016/j.jaci.2010.07.029
- Rosati, E., Dowds, C. M., Liaskou, E., Henriksen, E. K. K., Karlsen, T. H., & Franke, A. (2017). Overview of methodologies for T-cell receptor repertoire analysis. BMC Biotechnology, 17(1), 61. https://doi.org/10.1186/s12896-017-0379-9
- Sadelain, M., Brentjens, R., & Rivière, I. (2013). The basic principles of chimeric antigen receptor design. Cancer Discovery, 3(4), 388–398. https://doi.org/10.1158/2159-8290.CD-12-0548
- Schultz, L., & Mackall, C. (2019). Driving CAR T cell translation forward. Science Translational Medicine, 11(481). https://doi.org/10.1126/scitranslmed.aaw2127
- Seidel, U. J. E., Oliveira, C. C., Lampen, M. H., & Hall, T. van. (2012). A novel category of antigens enabling CTL immunity to tumor escape variants: Cinderella antigens. Cancer Immunology, Immunotherapy : CII, 61(1), 119–125. https://doi.org/10.1007/s00262-011-1160-x
- Sewell, A. K. (2012). Why must T cells be cross-reactive? Nature Reviews Immunology, 12(9), 669–677. https://doi.org/10.1038/nri3279
- Tang, X.-J., Sun, X.-Y., Huang, K.-M., Zhang, L., Yang, Z.-S., Zou, D.-D., Wang, B., Warnock, G. L., Dai, L.-J., & Luo, J. (2015). Therapeutic potential of CAR-T cell-derived exosomes: a cell-free modality for targeted cancer therapy. Oncotarget, 6(42), 44179–44190. https://doi.org/10.18632/oncotarget.6175
- Tran, K. Q., Zhou, J., Durflinger, K. H., Langhan, M. M., Shelton, T. E., Wunderlich, J. R., Robbins, P. F., Rosenberg, S. A., & Dudley, M. E. (2008). Minimally cultured tumor-infiltrating lymphocytes display optimal characteristics for adoptive cell therapy. Journal of Immunotherapy (Hagerstown, Md. : 1997), 31(8), 742–751. https://doi.org/10.1097/CJI.0b013e31818403d5
- Turner, S. J., Doherty, P. C., McCluskey, J., & Rossjohn, J. (2006). Structural determinants of T-cell receptor bias in immunity. Nature Reviews Immunology, 6(12), 883–894. https://doi.org/10.1038/nri1977
- Turul, T., Tezcan, I., Artac, H., de Bruin-Versteeg, S., Barendregt, B. H., Reisli, I., Sanal, O., van Dongen, J. J. M., & van der Burg, M. (2008). Clinical heterogeneity can hamper the diagnosis of patients with ZAP70 deficiency. European Journal of Pediatrics, 168(1), 87. https://doi.org/10.1007/s00431-008-0718-x
- Van Der Bruggen, P., Zhang, Y., Chaux, P., Stroobant, V., Panichelli, C., Schultz, E. S., Chapiro, J., Van den Eynde, B. J., Brasseur, F., & Boon, T. (2002). Tumor-specific shared antigenic peptides recognized by human T cells. Immunological Reviews, 188(1), 51–64. https://doi.org/10.1034/j.1600-065X.2002.18806.x
- Walseng, E., Wälchli, S., Fallang, L.-E., Yang, W., Vefferstad, A., Areffard, A., & Olweus, J. (2015). Soluble T-Cell Receptors Produced in Human Cells for Targeted Delivery. PLOS ONE, 10(4), e0119559. https://doi.org/10.1371/journal.pone.0119559
- William E. Paul. (2008). Fundamental Immunology. In Lippincott Williams & Wilkins; Sixth Edition edition (May 1, 2008) (Vol. 1).
- Yanagi, Y., Yoshikai, Y., Leggett, K., Clark, S. P., Aleksander, I., & Mak, T. W. (1984). A human T cell-specific cDNA clone encodes a protein having extensive homology to immunoglobulin chains. Nature, 308(5955), 145–149. https://doi.org/10.1038/308145a0