Lac Operon gene regulation; The lactose operon (Lac operon) is a group of genes that contains a single promoter which helps in encoding genes for transport and metabolism of lactose in E. coli and many other enteric bacteria. This transcriptional regulation mechanism of E. coli was studied by researchers Francois Jacob and Jacques Monod in 1961.
In prokaryotes, gene regulation is carried out by Lac Operon model in which E. coli and many other bacteria’s protein encoded genes are located in a single transcription unit called an operon which share the same transcriptional regulation but are translated individually. Physiological and environmental conditions influence the alteration in gene expression in prokaryotes.
Structure of Lac operon
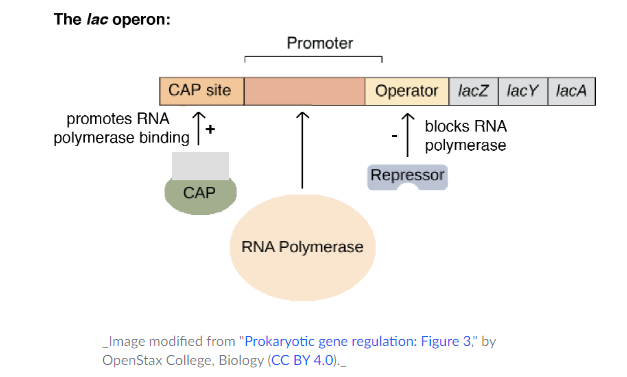
- The structural genes in the lac operon are the lacZ, lacY and lacA genes which are transcribed as a single mRNA in a single promoter.
- These lacZ, lacY and lacA gene specifies protein that help the cell utilize lactose and are encoded enzyme galactosidase, galactosidase permease and thiogalactosidase transacetylase respectively.
- Galactosidase permease transports lactose into the cell. It is also known as lactose permease.
- Galactosidase hydrolyzes lactose into glucose and galactose.
- Thiogalactosidase transacetylase transfers an acetyl group to galactosides, glucosides and lactosides.
- The enzyme that encodes lacZ splits lactose into monosaccharide that can be used in glycolysis.
- Similarly, lacY helps in membrane embedded transporter that brings lactose into the cell.
- The lac operon also contains a number of regulatory sequences of DNA where the proteins bind and control the transcription of the operon.
- Transcription occurs from a single operator where RNA polymerase binds.
- Between the promoter and the structural genes, there is operator which is a negative regulatory site bound by the lac repressor protein.
- When the lac repressor protein is bound, the operator overlaps the promoter which blocks the RNA polymerase.
- Catabolite activator protein (CAP) is bound in the positive regulatory site – CAP binding site that helps RNA polymerase bind to the promoter for further transcription.
What turns on lac operon?
In a media containing both lactose and glucose, E. coli prefers glucose over lactose as glucose requires less energy to break down as compare to lactose. But if lactose is only present in media, it utilizes it as an energy source. E. coli undergoes lac operon only when two conditions are favored:
- Availability of lactose and
- Glucose not available
To utilize lactose as a sugar source, E. coli must express its lac operon genes for lactose uptake and metabolism.
Lac operon in the presence of lactose
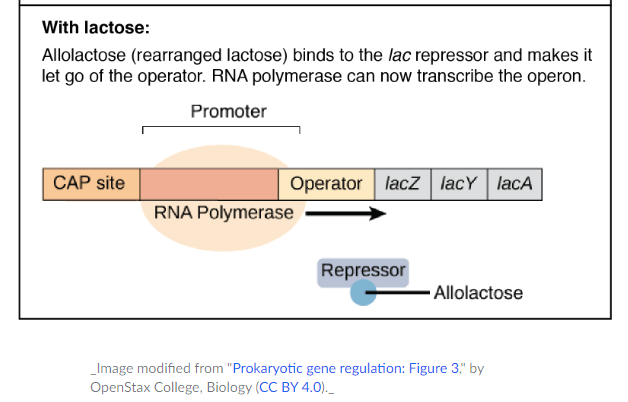
- When lactose is available, some molecules will be converted to allolactose, an isomer of lactose which binds to the lac repressor and changes its conformation to inhibit the binding to DNA.
- The lac repressor is the protein that inhibits the transcription in the lac operon which loses its ability to bind DNA and floats off the operator in presence of lactose.
- Allolactose is an inducer that turns on the lac operon which usually remains off.
- As it is usually off, it is also considered as an inducible operon.
Lac operon in the absence of lactose
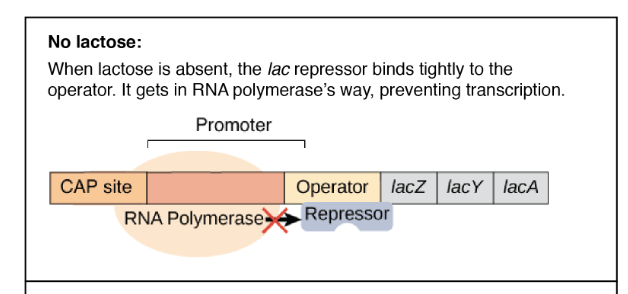
- When lactose is not present, the lac repressor binds tightly to the operator and prevents transcription of RNA polymerase and lacZ, lacY and lacA genes.
Lac operon in the presence of high glucose
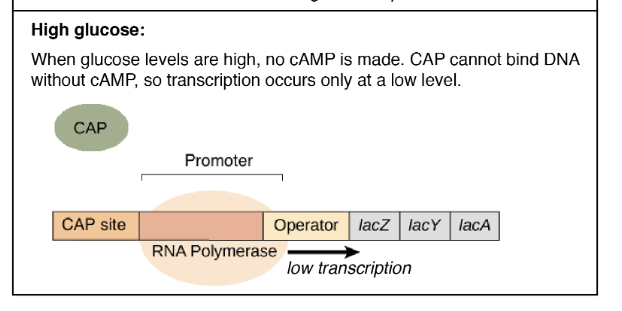
- When the glucose is present in the media, E. coli does not require lactose as a sugar source so lac operon does not need to be activated.
- Glucose inhibits the synthesis of cAMP from ATP as adenylate cyclase enzyme is inhibited by the presence of glucose.
- Therefore, cAMP is not produced where CAP cannot bind to the DNA without cAMP and remains inactive which results in low transcription.
- cAMP is a ‘hunger signal’ of E. coli when the glucose level is low.
Lac operon in the presence of low glucose
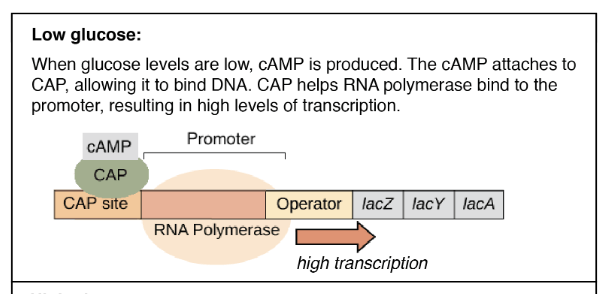
- When the glucose level is low, E. coli produces high level of cAMP which binds and activates the CRP (cAMP receptor protein), changes its conformation, binds to DNA and promotes transcription.
- When glucose is absent, cAMP binds to CRP that makes CRP-cAMP complex that stimulates the lac operon and allows the organism to utilize lactose as a carbon source.
- In the absence of glucose and presence of lactose, lac operon genes lacZ, lacY and lacA is highly transcribed.
How glucose and lactose levels are detected?
- Two regulatory proteins are involved in detecting the level of glucose and lactose in E. coli.
- The lac repressor acts as a lactose sensor and regulates its transcription based on the lactose level.
- The catabolite activator protein (CAP) functions as a glucose sensor that binds to the DNA of the lac operon and regulates its transcription based on the level of glucose present.
References:
- https://www.khanacademy.org/science/ap-biology/gene-expression-and-regulation/regulation-of-gene-expression-and-cell-specialization/a/the-lac-operon
- https://bio.libretexts.org/Bookshelves/Genetics/Book%3A_Online_Open_Genetics_(Nickle_and_Barrette-Ng)/12%3A_Regulation_of_Gene_Expression/12.01%3A_The_lac_Operon
- https://byjus.com/biology/lac-operon-regulation-gene-expression/
- https://courses.lumenlearning.com/boundless-biology/chapter/prokaryotic-gene-regulation/
- https://microbenotes.com/lac-operon/