Matrix-assisted laser desorption ionization-time of flight mass spectrometry (MALDI – TOF MS) is an emerging potential tool used for the identification and diagnosis of microorganisms. The use of MALDI – TOF MS is more as compared to other techniques because the process is rapid, has a high accuracy rate, and is economical in terms of cost and labor. MALDI – TOF MS can be used in broad areas like in the identification of bacteria, viruses, and certain fungal pathogens, strain typing, epidemiological studies, detection of water, food, and biological pathogens, detection of antibiotic-resistant microorganisms.
What is MALDI?
MALDI is “Matrix-Assisted Laser Desorption Ionization”. It is a soft ionization technique that strikes large molecules with laser energy into minimal ion fragments.
MALDI helps in the analysis of biomolecules such as DNA, proteins, peptides, saccharides, and large organic molecules like polymers, dendrimers, and other macromolecules. This technique is similar to electrospray ionization (ESI) as both contain soft ionization that helps in obtaining large ion molecules into gas-phase but MALDI produces much fewer multi ions than ESI.
MALDI: Principle and Methodology
(Source : https://www.creative-proteomics.com/technology/maldi-tof-mass-spectrometry.htm )
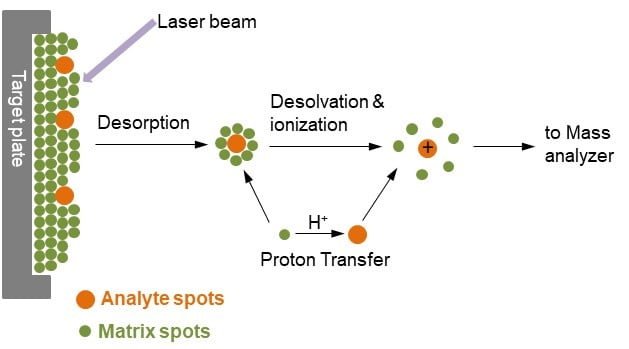
- The sample to be analyzed is mixed or coated with an organic compound that contains an absorbent solution called matrix. MALDI matrix functions as an energy absorbent upon laser irradiation. Commonly used matrixes are α-cyano-4-hydroxycinnamic acid, 2, 5-dihydroxybenzoic acid, 3, 5-dimethoxy-4-hydroxycinnamic acid, and 2, 6-dihydroxyacetophenone.
- The matrix compounds with the analyte are deposited on a target plate which is made up of a conducting metal. Then with a laser beam such as ultraviolet (UV) and infrared (IR), the sample is ionized on the spot where it becomes vibrationally excited.
- The matrix molecules energetically desorb and ionize in the presence of a laser beam and generate the analyte molecules into the gas phase.
- The protonated ion accelerates and gets separated on the basis of their mass and charge ratio. Small-sized ion molecules accelerate faster than large-sized ions.
- Then these protonated ions are detected and measured by time of flight (TOF) analyzer.
Time of flight (TOF) analyzer: Principle and methodology
(Source: https://www.creative-proteomics.com/technology/maldi-tof-mass-spectrometry.html )
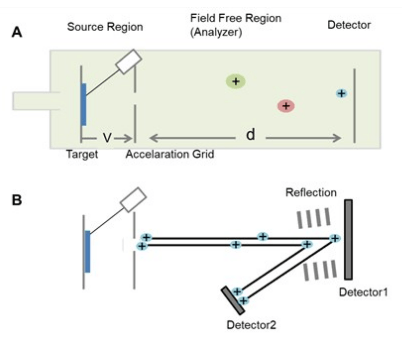
- The basic principle of the TOF analyzer is that it determines the time required by the ion of different size and charge ratio that takes to travel the length of the flight tube.
- The ion starts traveling at the same time where the small and lighter sized ions travel faster to the detector than the larger and heavier one.
- With the only use of liner TOF analyzer, all ions do not feel the pulse with equal intensity therefore, the reflector is applied. The reflector contains a high voltage series of ring electrodes and consists of a slightly displaced angle that helps the ion to pulse back in the flight tube.
- The ions that travel faster will spend more time within the reflector where the detector receives similar mass ions at the same time.
- TOF analyzer is high in resolution than other analyzers and is applicable only for microbiological purposes.
Mass spectrometry

(Source: https://www.creative-proteomics.com/technology/maldi-tof-mass-spectrometry.htm )
- The microorganisms are identified by comparing the masses of unknown organisms’ biomarkers with the proteome database.
- The probability mass function (PMF) is compared between the MS spectrum of unknown isolates and MS spectra of known isolates contained in the database.
- First, the analyte should be dissolved in a solvent making up to 0.1mg/ml and the matrix should be dissolved with a saturated or concentrated solution of about 10mg/ml. Both the solution is then mixed together in 1000:1 to 100,000:1 ratio.
- The mixture is placed on a metal target plate which crystallizes on drying and forms a solid deposit. Then the mixture is transferred into the MALDI – TOF instrument for analysis.
- Sublimation and ionization separate the ions depending upon the size and charge ratio through a TOF analyzer which is operated on the MS software.
- To increase the ability in identifying gram-positive and sugar non fermenting bacterial species, formic acid is used with the preparatory extraction of microbes whereas gram-negative bacteria can be identified using direct cell profiling.
Application of MALDI – TOF Mass Spectrometry:
Biochemistry:
MALDI – TOF MS is used in the field of biochemistry to identify the proteins and characterize it because protein contains a molecular weight that has an intact structure that becomes fragmented when ionized.
Peptide mass fingerprinting (PMF):
Peptide mass fingerprinting is used in the identification of proteins from simple mixture where MALDI – TOF plays an important role by its simple operation with high resolution and good mass accuracy. Peptides are generated by enzyme-like trypsin and are analyzed by MALDI – TOF MS and the obtained peptide masses are compared with the database.
Clinical and Environmental Bacteriology:
For a rapid, reliable, and cost-effective diagnosis of clinical and environmental samples, MALDI – TOF MS is appropriate for the early identification of bacterial culture. MALDI – TOF can detect blood culture, stool and urine samples, cerebrospinal fluids, and respiratory tract infections.
It can also identify food and waterborne pathogens and environmental samples.
Detection of viruses:
Besides immunological methods, researchers have proved using MALDI – TOF MS as more advanced and effective in use for diagnosing infectious viruses like influenza viruses, herpes viruses, and hepatitis viruses.
First, the viral genetic materials are amplified by PCR and later identified by MALDI – TOF MS which gives rapid and reliable results in a short period.
Organic chemistry:
Some synthetic macromolecules with a high molecular weight such as catenanes, rotaxanes, and hyperbranched polymers can be analyzed rapidly with effective results.
Medicine:
MALDI – TOF MS may even serve as an early detection technique for various types of cancer like it has been used in identifying membrane protein associated with pancreatic cancer.
It can also be used in determining drug resistance bacteria which could help physicians to decide before prescribing the medicine.
Learn more
References:
- https://www.frontiersin.org/articles/10.3389/fmicb.2015.00791/full
- https://en.wikipedia.org/wiki/Matrix-assisted_laser_desorption/ionization
- https://www.creative-proteomics.com/technology/maldi-tof-mass-spectrometry.htm
- https://journals.lww.com/oncology-times/fulltext/2003/10100/maldi_tof_mass_spectrometry__getting_a_feel_for.11.aspx
- https://academic.oup.com/femsre/article/36/2/380/565595