What is CRISPR?
CRISPR (Clusters of Regularly Interspaced Short Palindromic Repeats) technology is an emerging and powerful tool for editing genomes which allows researchers to easily alter DNA sequences and modify gene function used widely in correcting genetic defects, treating and preventing the spread of diseases and improving crops.
“CRISPR” is shorthand for “CRISPR-Cas9.” CRISPRs are specialized stretches of DNA clustered regularly interspaced short palindromic repeats acting like a pair of molecular scissors. They belong to the family of DNA sequences found in genome of prokaryotes and archaea. They are known to be derived from DNA fragments of bacteriophages that infected the prokaryote as the natural defense mechanisms. It was discovered that bacteria transcribe this DNA to RNA upon viral infection so that RNA is guided with a protein, Cas,” that cleaves DNA, providing protection against the virus.
How does CRISPR work?
The CRISPR immune system works through three basic steps:
- Adaption of CRISPR,
- Processing of CRISPR RNA
- Targeting with Cas9 protein
1. Adaption of CRISPR:
The specialized region of DNA of CRISPR has two distinct regions; the presence of nucleotide repeats and spacers.
- Repeated sequences of nucleotides — the building blocks of DNA — are distributed throughout a CRISPR region.
- Spacers are bits of DNA that are interspersed among these repeated sequences.
In bacteria, the spacers are indicated from viruses that attacked the organisms. Therefore, they serve as a bank of memories, which enables bacteria to recognize the viruses and fight off future attacks. This was first demonstrated experimentally by Rodolphe Barrangou and a team of researchers at Danisco, in 2007 published in the journal Science, the researchers used Streptococcus thermophilus bacteria, in yogurt and other dairy cultures, as their model. Thus, it was concluded that, new spacers were incorporated into the CRISPR region after viral attack.
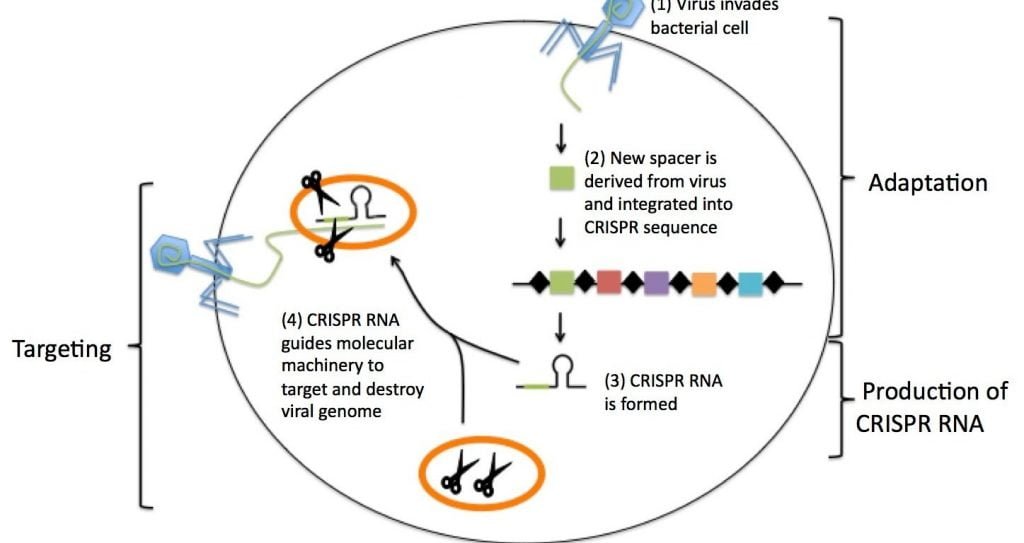
[Source; Barrangou, R. and Marraffini, L. (2014). CRISPR-Cas Systems: Prokaryotes Upgrade to Adaptive Immunity Molecular Cell 54, 234-244].
2. Processing of CRISPR RNA (crRNA)
When a spacer is incorporated after virus attack a portion of the CRISPR is transcribed and processed into CRISPR RNA, or “crRNA” where the nucleotide sequence of the CRISPR acts as a template to produce a complementary sequence of single-stranded RNA. The crRNA also consists of spacer and the repetitive sequences.
3. Targeting with Cas9 protein
The Cas9 protein is an enzyme that is capable of cleaving foreign DNA. The protein typically binds to two RNA molecules: crRNA and another called tracrRNA (or “trans-activating crRNA”). Thus, the crRNA guides Cas9 to the target site where it will make its cut. The target DNA is complementary to the 20-nucleotide stretch of crRNA.
Using two separate regions, or “domains” on its structure, Cas9 cuts both strands of the DNA double helix, making what is known as a “double-stranded break,”.
The built-in safety mechanism, which ensures that Cas9 doesn’t just cut anywhere in a genome is the short DNA sequences known as PAMs (“protospacer adjacent motifs”) that serve as tags and sit adjacent to the target DNA sequence. If the Cas9 complex doesn’t see a PAM next to its target DNA sequence, it won’t cut. Therefore, it is not possible to attack by Cas9 to attack the DNA sequence.
Application of CRISPR
With the advancement in technology, CRISPR genome editing allows scientists to quickly create cell and animal models, which researchers can use to accelerate research into medicine, treatments, agriculture, food industries and now being developed as a rapid diagnostic. Some of the major applications are as follows.
Treatment of genetic diseases
Research used this technology to treat genetic defect such as in mice, suggesting it could one day be used to treat the same type of hearing loss in people. These findings suggests that protein–RNA complex delivery of target gene-disrupting agents in vivo is a potential strategy for the treatment of some types of autosomal-dominant hearing loss. Another research is using it on mutations that cause Huntington’s disease or cystic fibrosis and trying it on the BRCA-1 and 2 mutations linked to breast and ovarian cancers. Scientists have even shown that CRISPR can knock HIV infections from T cells.
ii. Agriculture:
The genome editing tool is also used in agriculture for the production of transgenic crop to the select the desired variety crops. A researcher at Penn State University, used a popular gene editing tool called CRISPR/Cas9 to snip out a tiny piece of DNA from one particular gene in a white button mushroom thereby disabling the gene, which in turn reduces the mushroom’s production of an enzyme called polyphenol oxidase. This results in delaying of, the mushroom to turn brown. Recently, major companies like Monsanto and DuPont have begun licensing CRISPR technology, hoping to develop valuable new crop varieties with desired crop traits.
iii. Gene drives of the species
CRISPR technology can be also used for pest control. One of the recent examples is the gene drives for malaria. It is able to wipe out entire populations of malaria-spreading mosquitoes or resurrect once-extinct species like the passenger pigeon in the future.
By harnessing this technique, genetically modify mosquitoes to only produce male offspring — and then use a gene drive to push that trait through an entire population. Over time, the population would go extinct.
iv. Medicine:
The emergence of antimicrobial resistance demands an alternative in the global scenario. CRISPR are used in treatment of infectious diseases. They provide a way to make more specific antibiotics and antivirals that target only disease-causing strains and sparing beneficial bacteria. There are also researches on CRISPR systems that target viruses such as HIV and herpes. A recent advance in research also shows with the white blood cells defense improvement with HIV.
v. Modification of disease in babies:
One of the major research projects done was by a scientist in China, He Jiankui. He created the world’s first human babies with CRISPR-edited genes: a pair of twin girls resistant to HIV.
vi. Food Industry
It is used in food industries to produce allergy free food, minimize the food-toxins, improve strains. With CRISPR, it could be possible to make milk, eggs or peanuts allergy free. Using CRISPR, the company has been able to turn off the genes that make the beans produce caffeine. In fact, the original discovery of CRISPR immunity came from researchers at Danisco, from S. thermophilus with immunity against such viral attack.
vii. Laboratory Analysis and Research
Beyond applications encompassing bacterial immune defenses, scientists can use this technology as a convenient technique in the lab to make precise changes in the genes of organisms as diverse as fruit flies, fish, mice, plants and even human cells for their study.
Major findings on CRISPR
There have been many recent research projects based around CRISPR which is developing CRISPR-based solutions for medicine, agriculture, and biological research.
- In 2012, there were two research papers were published in the Journals Science and PNAS, which helped transform bacterial CRISPR-Cas9 into a simple, programmable genome-editing tool. This was very important discovery for establishing CRISPR-Cas as genome editing tool. Another similar studies, done by separate groups, concluded that Cas9 could be directed to cut any region of DNA which was performed by changing the nucleotide sequence of crRNA, which binds to a complementary DNA target.
- In the 2012 Science article, Martin Jinek and colleagues further demonstrated and simplified the system by fusing crRNA and tracrRNA to create a single “guide RNA” which concluded genome editing requires two components: a guide RNA and the Cas9 protein.
- On November 2017, in the Journal Nature Communications, a team of researchers led by Kanazawa University and University of Tokyo showed the mechanism of CRISPR is in action for the very first time.
- On April 2017, as released research in the Journal Science a CRISPR molecule was programmed to find strains of viruses, such as Zika, in blood serum, urine and saliva.
- On June 2017 as published on Nature, (CRISPR)/CRISPR-associated nuclease 9 (Cas9) system is an emerging gene-editing technique with the potential to eliminate or disrupt HIV-integrated genomes or HIV-infected cells from multiple HIV reservoirs. Encouraging progress is made for HIV/AIDS treatment and prevention, both in vitro in human patient cells and in vivo in animal model experiments by this technology.
- On Aug. 2017, scientists revealed removal a heart disease defect in an embryo successfully using CRISPR as published on Nature.
- On January 2018, researchers announced that CRISPR to inhibit plant pathogen such ass fungi and other problems that threaten chocolate production making the plants variety resistant to disease.
- On November 2019 by CRISPR Therapeutics, Switzerland, and Vertex Pharmaceuticals company announcement of a CRISPR–Cas9 therapy clinical trial suggests potentially curative responses in two patients with transfusion-dependent β-thalassemia and sickle cell disease, respectively.
- In September 2019, a company used this tool for crops that are more productive and consume less water and fertilizer than those currently produced by seed conglomerates. Inari is one of a several small companies with similarly lofty goals who are capitalizing on new editing technologies, such as CRISPR, and computational methods for predictive modeling makes crop development faster and less expensive providing opportunity for startups.
- Recent advances in research in 2020 showed that CRISPR/Cas9 has been applied in generating animal models as well as gene therapy in vivo of retinitis pigmentosa (RP) and Leber congenital amaurosis (LCA). Thus, the CRISPR technology is a potential attempt for clinical trial with combination of other technologies such as adeno-associated virus (AAV) and induced pluripotent stem cells (iPSCs).
CD4 T Lymphocyte Cells: Types, Functions, CD4 Count and HIV (thesciencenotes.com)
References
- Barrangou R (2015). “The roles of CRISPR-Cas systems in adaptive immunity and beyond.” Current Opinion in Immunology. 32: 36–41. doi:10.1016/j.coi.2014.12.008.
- Barrangou, R. and Marraffini, L. (2014). “CRISPR-Cas Systems: Prokaryotes Upgrade to Adaptive Immunity.” Molecular Cell, 54, 234-244.
- Doudna J.A., Charpentier E. (2014). “The new frontier of genome engineering with CRISPR-Cas9.” Science, Vol. 346, Issue 6213, 1258096. DOI: 10.1126/science.1258096.
- Gao, X., Tao, Y., Lamas, V., et al. (2018). “Treatment of autosomal dominant hearing loss by in vivo delivery of genome editing agents.” Nature, 553, 217–221. https://doi.org/10.1038/nature25164.
- Hille F, Richter H, Wong SP, Bratovič M, Ressel S, Charpentier E (2018). “The Biology of CRISPR-Cas: Backward and Forward.” Cell, 172(6): 1239–1259. doi:10.1016/j.cell.2017.11.032.
- Huang, Z., Tomitaka, A., Raymond, A., et al. (2017). “Current application of CRISPR/Cas9 gene-editing technique to eradication of HIV/AIDS.” Gene Therapy, 24, 377–384. https://doi.org/10.1038/gt.2017.35.
- Jinek, M., Chylinski, K., Fonfara, I., Hauer, M., Doudna, J. A., & Charpentier, E. (2012). “A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity.” Science (New York, N.Y.), 337(6096), 816–821. https://doi.org/10.1126/science.1225829.
- Mulepati S, Héroux A, Bailey S (2014). “Crystal structure of a CRISPR RNA–guided surveillance complex bound to a ssDNA target.” Science, 345(6203): 1479–1484. doi:10.1126/science.1256996.
- “Nature Biotechnology” 37, 1251-1252 (2019). doi: 10.1038/d41587-019-00027-2.
- Osdaghi E., Martins S.J., Ramos-Sepulveda L., Vieira F.R., Pecchia J.A., Beyer D.M., Bell T.H., Bull C.T. (2019). “100 years since Tolaas: Bacterial blotch of mushrooms in the 21st century.” Plant Disease, 103(11), pp. 2714-2732.
- Peng, Y. Q., Tang, L. S., Yoshida, S., & Zhou, Y. D. (2017). “Applications of CRISPR/Cas9 in retinal degenerative diseases.” International Journal of Ophthalmology, 10(4), 646–651. https://doi.org/10.18240/ijo.2017.04.23.
- Pennisi, E. (2013) “The CRISPR Craze.” Science, 341(6148), 833-836.
- Sander, J., Joung, J. (2014). “CRISPR-Cas systems for editing, regulating and targeting genomes.” Nature Biotechnology, 32, 347–355. https://doi.org/10.1038/nbt.2842.