Enzyme-linked immunosorbent assay (ELISA) is a versatile and highly sensitive technique used to measure the presence and concentration of a wide variety of biomolecules, including antigens, antibodies, peptides, proteins, and hormones, in biological samples. The sensitivity of ELISA arises from its ability to detect interactions between a single antigen-antibody complex, combined with enzymatic amplification, which converts a colorless substrate into a chromogenic or fluorescent product measurable with a microplate reader. ELISA is broadly classified into four main types—direct, indirect, sandwich, and competitive—each optimized for specific experimental needs, sample types, and available antibodies.
Principles of ELISA
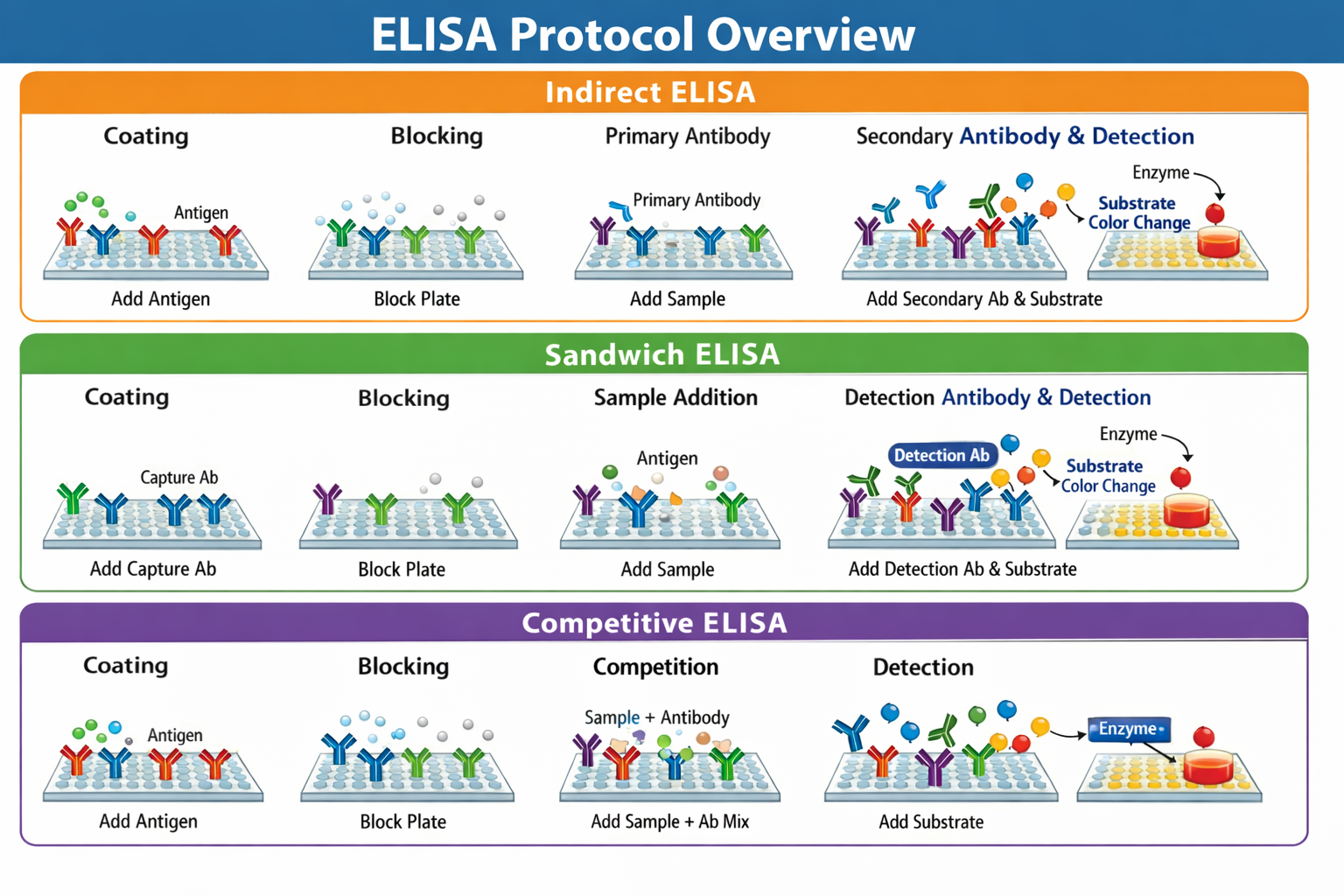
All ELISA methods share the common principle of immobilizing a target analyte on a solid surface, detecting it using specific antibodies or antigens, and generating a measurable signal through an enzyme-catalyzed reaction. In direct ELISA, the target antigen is directly detected by an enzyme-conjugated antibody. Indirect ELISA uses a secondary enzyme-conjugated antibody to detect a primary antibody bound to an immobilized antigen. Sandwich ELISA captures the antigen between two specific antibodies—one immobilized as a capture antibody and one enzyme-conjugated detection antibody. Competitive ELISA measures antigen concentration based on the competition between a sample antigen and plate-bound antigen for a limited amount of antibody.
Common enzymes used include horseradish peroxidase (HRP) and alkaline phosphatase (AP), which convert colorless substrates such as 3,3′,5,5′-tetramethylbenzidine (TMB), 2,2’-azino-bis(3-ethylbenzothiazoline-6-sulfonic acid) (ABTS), or p-nitrophenyl phosphate (PNPP) into detectable chromogenic products. The choice of substrate, enzyme, and plate reader wavelength is crucial for optimizing sensitivity and signal detection.
Indirect ELISA Protocol
Purpose: Quantify antibodies in serum, hybridoma supernatants, or other biological fluids.
Materials
96-well flat-bottom ELISA plate
Purified antigen (e.g., 2 mg/mL Influenza A virus)
PBS (1X), pH 7.4
Tween-20 (1%) in PBS
Blocking buffer: 5% donkey serum in PBS
Primary antibody (sample containing antibody)
Secondary enzyme-conjugated antibody (HRP or AP)
Substrate (TMB for HRP, PNPP for AP)
Stop solution (2N H₂SO₄)
Procedure
Coating the Plate: Pipette 50 µL of purified antigen into each well. Cover the plate and incubate overnight at 4°C for antigen adsorption. Remove unbound antigen by flicking the plate over a sink.
Blocking: Add 200 µL blocking buffer per well to prevent non-specific binding. Incubate 2 hours at room temperature or overnight at 4°C, then wash wells three times with PBS + 1% Tween-20.
Primary Antibody Incubation: Prepare serial dilutions of serum or sample (e.g., 1:12.5 to 1:204,800 in PBS). Add 100 µL of each dilution to the appropriate wells. Incubate 1–2 hours at room temperature or overnight at 4°C, then wash 3–5 times with PBS + 1% Tween-20.
Secondary Antibody Incubation: Add 100 µL of enzyme-conjugated secondary antibody (e.g., HRP-donkey anti-mouse IgG) per well. Incubate 1 hour at room temperature, then wash 3–5 times.
Detection: Add 100 µL substrate (TMB 1 mg/mL) to each well. Incubate 5–10 minutes, then stop the reaction with 100 µL 2N H₂SO₄. Read absorbance at 450 nm (TMB) or 405 nm (PNPP).
Notes: Mean optical density (OD) values from triplicates are used to generate standard curves or semi-quantitative analysis.
Sandwich ELISA Protocol
Purpose: Measure antigens in complex biological samples, including cytokines and tissue lysates.
Materials
96-well ELISA plate
Capture antibody (1–10 µg/mL)
Detection antibody (enzyme-conjugated)
Sample containing antigen
Blocking buffer (5% nonfat dry milk in PBS)
PBS + 1% Tween-20
Substrate and stop solution
Procedure
Coating the Plate: Add 100 µL of capture antibody per well. Cover and incubate overnight at 4°C. Remove unbound antibody by flicking the plate.
Blocking: Add 200 µL blocking buffer (5% nonfat dry milk in PBS) per well. Incubate 2 hours at room temperature, then wash 3–5 times with PBS + 1% Tween-20.
Adding Samples: Pipette 100 µL of sample containing antigen into each well. Seal and incubate 1–2 hours at room temperature or overnight at 4°C. Wash 3–5 times.
Detection Antibody: Add 100 µL enzyme-conjugated detection antibody to each well. Incubate 2 hours at room temperature, then wash 3–5 times.
Detection: Add 100 µL substrate per well. Incubate 5–10 minutes at room temperature. Stop reaction with 100 µL 2N H₂SO₄. Read absorbance at 450 nm.
Notes: Standard curves prepared from known antigen concentrations are required for quantitative analysis.
Competitive ELISA Protocol
Purpose: Quantify small antigens or antigens with a single available antibody.
Materials
96-well ELISA plate
Purified antigen (1–10 µg/mL) for coating
Primary antibody (enzyme-conjugated or unconjugated)
Sample antigen
Secondary antibody (if needed)
Blocking buffer
Substrate and stop solution
Procedure
Plate Coating: Add 100 µL purified antigen per well. Cover and incubate overnight at 4°C. Remove unbound antigen.
Blocking: Add 200 µL blocking buffer per well. Incubate 2 hours at room temperature, then wash 3–5 times with PBS + 1% Tween-20.
Antigen-Antibody Pre-incubation: Mix 150 µL sample antigen with 150 µL primary antibody in a separate tube. Incubate 1 hour at 37°C.
Competition: Add 100 µL of antigen-antibody mixture to wells. Incubate 1 hour at 37°C, then wash 3–5 times.
Secondary Antibody (if needed): Add 100 µL enzyme-conjugated secondary antibody. Incubate 1 hour, then wash 3–5 times.
Detection: Add 100 µL substrate per well. Incubate 5–10 minutes, stop with 100 µL 2N H₂SO₄, and read absorbance.
Notes: In competitive ELISA, higher antigen concentrations produce lower signals.
Data Analysis
For both indirect and sandwich ELISAs, mean OD values from replicates are plotted against sample dilutions or known standards to generate standard curves. Absorbance values from unknown samples are compared to these curves to determine analyte concentrations. It is critical to ensure that measured values fall within the linear range of the standard curve. For indirect ELISA, serial dilutions of serum or antibodies ensure precise quantification, while sandwich ELISA allows sensitive detection of antigen in complex samples. Competitive ELISA uses inverse correlation between signal and analyte concentration for quantification.
Summary of Strengths and Limitations
| ELISA Type | Strengths | Limitations |
|---|---|---|
| Indirect | High sensitivity; multiple secondary antibodies increase signal; flexible for different primary antibodies | High background; requires well-coated antigen |
| Sandwich | Highly specific and sensitive; works with complex samples | Requires optimization of antibody pairs; limited to analytes with multiple epitopes |
| Competitive | Can use unpurified samples; suitable for small molecules | Requires large amounts of pure antigen; less sensitive to reagent dilution |
ELISA remains a cornerstone of immunology and biochemistry research due to its sensitivity, versatility, and quantitative capabilities. With careful adherence to protocol details—antigen and antibody concentrations, incubation times, washing steps, and substrate selection—ELISA provides reliable measurement of proteins, antibodies, and other analytes in diverse experimental settings.