Human macrophages are central players in innate immunity, inflammation, tissue repair, infection response, cancer biology, and neurodegenerative disease. Because they respond rapidly to environmental signals and help coordinate immune activity, macrophages are often used to study disease mechanisms that involve chronic inflammation, pathogen sensing, immune regulation, and cellular stress.
However, macrophage research has a persistent challenge: obtaining enough consistent, experimentally tractable human macrophages for large-scale functional studies. Primary macrophages derived from blood monocytes are useful, but they can vary substantially between donors and preparations. This variability can complicate reproducibility, especially in pooled genetic screens or high-throughput functional genomics experiments.
Human induced pluripotent stem cell-derived macrophages, often abbreviated as iPSC-derived macrophages, offer a powerful alternative. They can be generated from pluripotent stem cell systems, expanded through controlled differentiation workflows, and used to model macrophage biology in a more standardized way. When combined with CRISPR-Cas9 gene knockout technology, these cells become a valuable platform for identifying genes that regulate inflammation, innate immune activation, host-pathogen interactions, and disease-relevant macrophage phenotypes.
A 2024 STAR Protocols article by Navarro-Guerrero and colleagues describes an optimized approach for CRISPR-Cas9 knockout in human iPSC-derived macrophages using lentiviral delivery supported by VPX virus-like particles. The protocol focuses on practical issues that often limit CRISPR screening in macrophage-like cells, including lentiviral transduction efficiency, antibiotic selection, library titration, and cell viability.
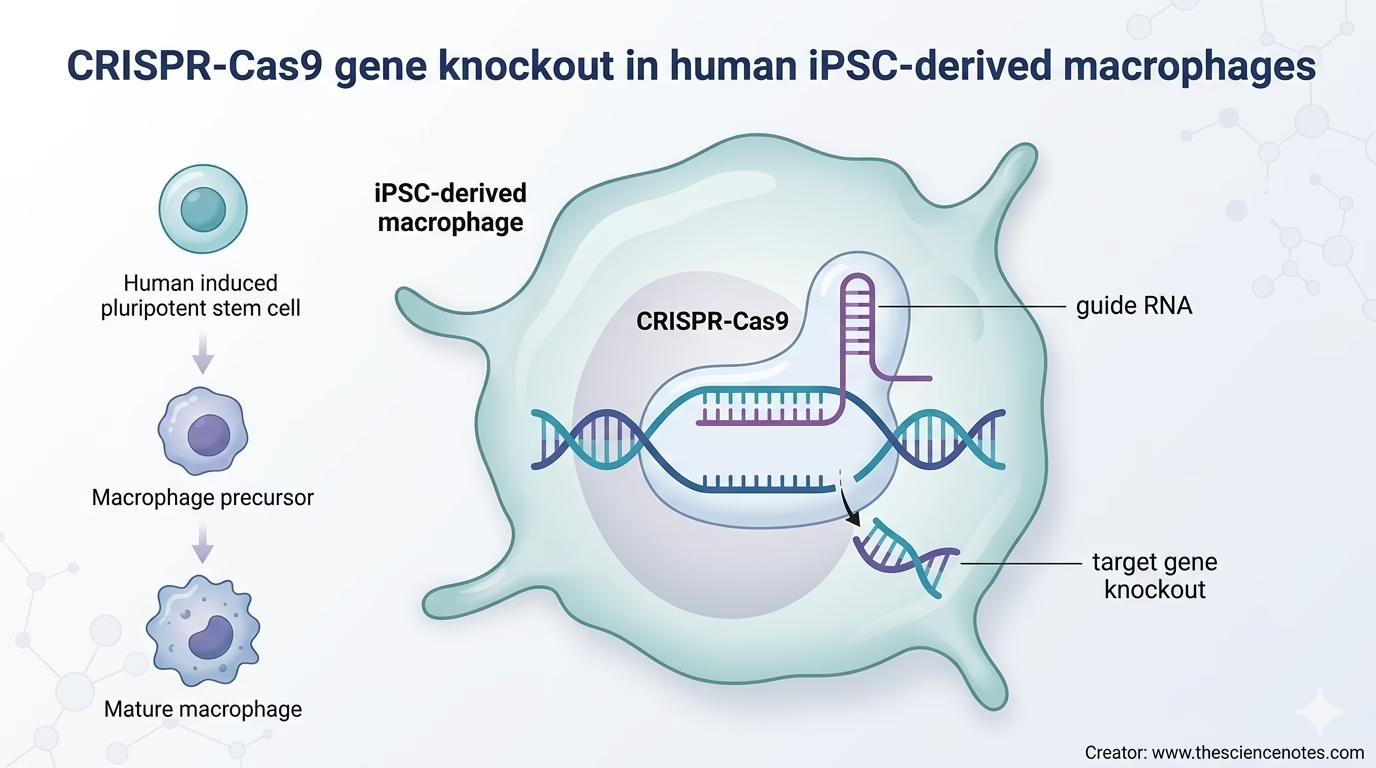
What Are iPSC-Derived Macrophages?
iPSC-derived macrophages are macrophage-like immune cells produced from induced pluripotent stem cells. Induced pluripotent stem cells are adult cells that have been reprogrammed into a pluripotent state, allowing them to differentiate into many cell types under controlled conditions.
In macrophage research, iPSC-derived systems are attractive because they can provide a more scalable and consistent source of human macrophages than traditional primary monocyte-derived macrophage models. This is especially valuable for genome-wide CRISPR screens, where researchers may need large numbers of cells while maintaining consistent differentiation status and phenotype.
A key requirement for successful CRISPR screening in these cells is the generation of a homogeneous macrophage population. The STAR Protocols article emphasizes the importance of confirming macrophage identity and immune responsiveness before screening. Markers such as CD11b and CD14 are commonly used to support macrophage characterization, while functional activation can be assessed through inflammatory response markers after stimulation.
In practical terms, the better the macrophage differentiation, the more interpretable the CRISPR results. Poorly differentiated cells may produce misleading phenotypes, reduce screening sensitivity, or obscure the role of genes involved in macrophage-specific biology.
Why Use CRISPR-Cas9 Knockout in iPSC-Derived Macrophages?
CRISPR-Cas9 knockout technology allows researchers to disrupt specific genes and observe the resulting changes in cell behavior. In pooled CRISPR screening, thousands of guide RNAs can be introduced into a cell population, allowing systematic identification of genes that influence survival, activation, cytokine production, infection response, or other phenotypes.
For macrophages, this approach is especially useful because many disease-relevant pathways involve complex regulatory networks. These include:
- Pattern recognition receptor signaling
- Cytokine production
- Phagocytosis
- Antigen presentation
- Inflammasome activation
- Lipid handling
- Host-pathogen interactions
- Immune suppression in cancer
- Neuroinflammatory signaling
By knocking out genes across the genome or within targeted gene sets, researchers can identify regulators that would be difficult to discover using candidate-gene approaches alone.
The TKOv3 CRISPR library discussed in the source protocol is an example of a pooled genome-wide knockout library. It targets thousands of human protein-coding genes using multiple guide RNAs per gene, making it suitable for functional genomics experiments where comprehensive gene interrogation is needed. Its relatively compact design is also useful when cell numbers are limited, which is often the case in iPSC-derived immune cell systems.
The Challenge: Delivering CRISPR Components into Macrophages
Macrophages are not always easy to genetically manipulate. Lentiviral vectors are commonly used to deliver CRISPR-Cas9 systems into mammalian cells, but macrophages and related myeloid cells can restrict viral transduction. This can reduce editing efficiency and compromise pooled screening quality.
One reason for this difficulty involves antiviral restriction factors. SAMHD1, for example, is known to restrict lentiviral transduction in certain immune cells by limiting the intracellular environment needed for efficient reverse transcription. The STAR Protocols workflow uses VPX virus-like particles to help overcome this restriction.
VPX is a viral accessory protein associated with degradation of SAMHD1. In the protocol, VPX virus-like particles are used alongside lentiviral CRISPR delivery to improve transduction efficiency in iPSC-derived macrophages. Western blotting for SAMHD1 is used as a quality-control readout to assess the effectiveness of VPX-VLP treatment.
This optimization is important because low transduction efficiency can distort pooled CRISPR screens. If too few cells receive guide RNAs, the screen may lose library representation. If too many cells receive multiple guide RNAs, gene-level interpretation becomes more complicated. A carefully optimized transduction workflow helps preserve both biological signal and screening integrity.
Core Elements of an Optimized CRISPR Knockout Workflow
Although the published protocol contains detailed laboratory steps, the overall workflow can be understood through several major phases.
1. Confirm Macrophage Differentiation and Function
Before performing CRISPR knockout or screening, researchers must ensure that the iPSC-derived cells have differentiated into a consistent macrophage population. This includes checking marker expression and confirming that the cells can respond to innate immune stimuli.
This step is not optional. If cells are heterogeneous, poorly differentiated, or functionally impaired, downstream CRISPR phenotypes may reflect differentiation artifacts rather than true gene function.
2. Optimize Antibiotic Selection
Many lentiviral CRISPR vectors include an antibiotic resistance marker. After transduction, antibiotic selection enriches for cells that received the vector. However, antibiotic exposure can affect macrophage biology, viability, and inflammatory activation.
The protocol uses a viability-based approach to identify the minimum antibiotic concentration needed to eliminate non-transduced cells. This optimization helps balance selection stringency with preservation of macrophage health.
The source article also notes an important biological limitation: puromycin selection may influence immune activation. In the reported system, puromycin-treated transduced macrophages showed reduced TNF expression after LPS stimulation compared with controls. This observation matters because selection conditions can alter the phenotype being studied.
3. Determine Polybrene Tolerance
Polybrene is often used to improve lentiviral transduction by reducing charge-based repulsion between viral particles and the cell surface. However, it can also affect cell viability depending on concentration and cell type.
An optimized workflow therefore includes determining the highest polybrene exposure that improves transduction without compromising macrophage survival or function. This is especially important in sensitive or non-dividing immune cell models.
4. Produce and Titrate VPX Virus-Like Particles
VPX-VLPs are included to enhance lentiviral transduction in macrophage-like cells. The source protocol describes large-scale VPX-VLP production and subsequent titration using SAMHD1 depletion as a readout.
The key concept is that VPX-VLPs must be functional and appropriately matched to the target cell system. Too little VPX activity may fail to improve transduction. Poor-quality preparations may create inconsistent results. Excessive or poorly controlled exposure may affect cell health.
5. Determine Lentiviral CRISPR Library MOI
Multiplicity of infection, or MOI, refers to the average number of viral particles delivered per cell. In pooled CRISPR knockout screens, researchers often aim for low MOI so that most transduced cells receive only one guide RNA. This supports clearer assignment of phenotype to gene knockout.
The protocol uses antibiotic selection and viability measurement to determine the amount of lentiviral CRISPR library that produces the desired transduction range under the exact screening conditions. This is important because MOI cannot always be reliably transferred across plate formats, cell densities, media conditions, or viral preparations.
6. Perform CRISPR-Cas9 Knockout and Phenotypic Readout
Once the system is optimized, iPSC-derived macrophage precursors can be transduced with CRISPR-Cas9 lentiviral constructs. After differentiation and appropriate selection or validation, cells can be analyzed using viability assays, flow cytometry, cytokine readouts, protein detection, sequencing, or other phenotype-specific methods.
The endpoint depends on the research question. For example, a screen might identify genes that regulate inflammatory cytokine production, macrophage survival, phagocytosis, pathogen response, or disease-associated immune states.
Why VPX Virus-Like Particles Matter in Macrophage CRISPR Screening
The use of VPX-VLPs is one of the most important features of the optimized workflow. Macrophages and related myeloid cells can resist lentiviral transduction because of intrinsic antiviral mechanisms. VPX-VLPs help reduce this barrier by targeting SAMHD1, thereby improving the conditions for lentiviral delivery.
For CRISPR screening, this improvement can have several benefits:
Higher transduction efficiency: More cells receive CRISPR components, improving the feasibility of screens in cell-limited systems.
Better library representation: Sufficient representation of guide RNAs is critical for pooled screening accuracy.
Lower pressure to use excessive viral input: Better delivery can reduce the need for overly aggressive transduction conditions that may harm cells.
Improved reproducibility: Optimized VPX-VLP titration can reduce batch-to-batch variability.
However, VPX-VLP use also introduces another variable that must be validated. The protocol uses SAMHD1 detection by western blotting as a functional quality-control measure, which helps confirm whether VPX-VLP preparations are performing as intended.
Applications in Inflammation and Innate Immunity Research
CRISPR knockout in iPSC-derived macrophages is particularly valuable for studying innate immune pathways. Macrophages are among the first immune cells to detect infection, tissue injury, and inflammatory signals. They produce cytokines, engulf particles, coordinate immune responses, and shape tissue microenvironments.
An optimized CRISPR platform can support research into:
Chronic Inflammation
Many diseases involve persistent macrophage activation or immune dysregulation. These include atherosclerosis, inflammatory bowel disease, rheumatoid arthritis, and metabolic inflammatory conditions. CRISPR screens can help identify genes that promote or suppress inflammatory signaling.
Neurodegenerative Disease
Macrophage-like cells and tissue-resident immune cells are implicated in neuroinflammation. While microglia are the resident macrophages of the central nervous system, iPSC-derived macrophage systems can still help investigate broader myeloid immune mechanisms relevant to neurodegenerative disease.
Cancer Progression
Tumor-associated macrophages can suppress anti-tumor immunity, promote angiogenesis, remodel tissue, and influence response to therapy. CRISPR knockout studies can identify macrophage genes that regulate immune suppression, tumor interaction, or inflammatory signaling in cancer contexts.
Infectious Disease
Macrophages interact with bacteria, viruses, fungi, and parasites. They can eliminate pathogens, but some pathogens exploit macrophage biology for survival or dissemination. CRISPR screening can identify host factors involved in susceptibility or resistance to infection.
Sepsis and Hyperinflammation
Macrophage cytokine responses are central to systemic inflammatory syndromes. Functional genomics can help identify regulators of excessive cytokine production, immune paralysis, or inflammatory resolution.
Quality Control Considerations
A successful CRISPR knockout experiment in iPSC-derived macrophages depends on more than simply delivering Cas9 and guide RNAs. Several quality-control checkpoints are necessary.
Macrophage Identity
Researchers should confirm that cells express expected macrophage markers and display appropriate functional behavior. A homogeneous starting population improves confidence in downstream phenotypes.
Transduction Performance
Lentiviral delivery should be assessed under the same conditions used for the actual experiment. Changes in media, plate format, cell density, differentiation stage, viral batch, or additives can affect transduction.
VPX-VLP Function
Because VPX-VLPs are included to improve delivery, their activity should be confirmed. SAMHD1 reduction is a relevant readout in the published workflow.
Cas9 Expression
Cas9 expression should be validated, particularly in knockout workflows where inefficient editing would reduce phenotype detection.
Knockout Confirmation
For targeted knockout experiments, protein-level validation can confirm whether gene disruption has produced the expected loss of target expression.
Guide RNA Distribution
In pooled screens, next-generation sequencing can be used to examine guide RNA representation. Poor representation can reduce screen quality and lead to false negatives or biased results.
Limitations of CRISPR Knockout in iPSC-Derived Macrophages
Despite its strengths, this approach has limitations.
Antibiotic Selection Can Alter Macrophage Phenotype
The source protocol highlights that puromycin selection can reduce TNF expression after inflammatory stimulation. This is an important warning for researchers studying cytokine responses. Selection conditions must be incorporated into experimental design and control selection.
Differentiation Quality Is Critical
If iPSC-derived macrophages are not homogeneous or functionally mature, screening results may be difficult to interpret. Differentiation variability can introduce noise or bias.
Transduction Conditions Require Optimization
Macrophage transduction depends on multiple interacting variables, including viral quality, cell density, polybrene tolerance, VPX-VLP activity, spinfection conditions, and cell state. Small changes can affect efficiency and viability.
Cell Viability Can Limit Screens
CRISPR screens require enough viable cells to preserve library complexity. If transduction or selection reduces viability too much, library coverage may suffer.
Knockout Biology May Be Context-Dependent
The function of a gene in macrophages may depend on differentiation state, stimulation condition, disease model, genetic background, or assay timing. CRISPR results should be validated in relevant secondary assays.
Troubleshooting Concepts
The source protocol includes troubleshooting guidance for common problems. At a high level, these issues fall into several categories.
Poor Macrophage Differentiation
If cells do not express expected markers or respond poorly to stimulation, the differentiation system should be reviewed. Media quality, reagent freshness, and differentiation timing can all influence macrophage identity.
Inefficient VPX-VLP Production
VPX-VLP quality can be affected by plasmid quality, transfection performance, cell health, contamination, and producer-cell passage history. Consistency in production conditions is essential.
Low Transduction Efficiency
Low efficiency may result from reduced viral titer, repeated freeze-thaw cycles, inadequate transduction enhancement, suboptimal cell density, or insufficient VPX-VLP activity. The source protocol notes that lentiviral titer can decline with freeze-thaw cycles, so aliquoting and storage practices matter.
Low Cell Viability
Cell death after transduction can reflect excessive stress, poor plating density, viral toxicity, reagent toxicity, or selection pressure. Improving cell handling and optimizing exposure conditions can help preserve viability.
Insufficient Protein for Validation
Western blotting can require substantial protein input. When sample availability is limited, more sensitive automated protein detection systems may be appropriate.
Research Impact of the Optimized Protocol
The major contribution of the Navarro-Guerrero et al. protocol is that it adapts CRISPR-Cas9 knockout screening to a challenging but highly relevant human immune cell model. iPSC-derived macrophages provide a scalable platform, while VPX-VLP-assisted lentiviral delivery helps overcome a key technical barrier.
This combination enables researchers to ask gene-function questions in macrophages more systematically. Instead of studying one candidate gene at a time, researchers can screen broad libraries and identify regulators of immune phenotypes in a human cellular context.
For disease biology, this is especially important. Macrophages are not passive immune cells. They shape inflammation, tissue remodeling, tumor progression, infection outcome, and immune resolution. Understanding the genes that control these processes can reveal therapeutic targets and improve disease modeling.
Best Practices for Interpreting CRISPR Screen Results
CRISPR screens generate hypotheses. They do not automatically prove mechanism. Strong interpretation requires validation.
Researchers should consider:
Guide-level consistency: Multiple guide RNAs targeting the same gene should ideally show concordant effects.
Independent validation: Candidate genes should be tested in follow-up experiments using separate guide RNAs or orthogonal approaches.
Phenotype specificity: A gene affecting cytokine output may also affect cell viability, differentiation, or stress response. These possibilities should be separated experimentally.
Selection effects: Antibiotic selection and transduction conditions can influence immune activation and should be included in controls.
Biological context: A gene hit under one stimulation condition may not behave the same way under another.
This careful interpretation is particularly important in macrophages, where activation states are dynamic and highly responsive to environmental conditions.
Frequently Asked Questions
What is the main purpose of CRISPR knockout in iPSC-derived macrophages?
The main purpose is to identify genes that regulate macrophage biology. This can include genes involved in inflammation, innate immune activation, cytokine production, pathogen response, survival, differentiation, and disease-associated immune phenotypes.
Why are iPSC-derived macrophages useful for CRISPR screening?
They provide a scalable and relatively standardized human macrophage model. This helps overcome some limitations of primary macrophages, such as donor variability and limited cell availability.
Why is lentiviral delivery challenging in macrophages?
Macrophages have intrinsic antiviral defense mechanisms that can restrict lentiviral transduction. These mechanisms can reduce the efficiency of CRISPR component delivery.
What role do VPX virus-like particles play?
VPX virus-like particles help improve lentiviral transduction in macrophage-like cells by counteracting SAMHD1-related restriction. This can improve CRISPR delivery efficiency and support better screening performance.
Why is MOI important in pooled CRISPR screens?
Low MOI helps ensure that most transduced cells receive a single guide RNA. This makes it easier to link a cellular phenotype to disruption of a specific gene.
Can antibiotic selection affect macrophage behavior?
Yes. The source protocol reports that puromycin selection can reduce TNF expression after LPS stimulation in transduced iPSC-derived macrophages. This means selection conditions must be carefully controlled and interpreted.
What are common readouts after CRISPR knockout?
Common readouts include cell viability, flow cytometry, cytokine expression, western blotting, sequencing-based guide representation, and phenotype-specific functional assays.
Is this approach only useful for genome-wide screens?
No. The same general optimization principles can support targeted knockout studies, pathway-focused CRISPR libraries, and single-gene functional validation.
Conclusion
CRISPR knockout in iPSC-derived macrophages is a powerful approach for studying human innate immunity and macrophage-driven disease mechanisms. By combining iPSC-derived macrophage models with optimized lentiviral CRISPR-Cas9 delivery, researchers can investigate gene function at scale in a relevant human immune cell context.
The STAR Protocols workflow by Navarro-Guerrero and colleagues highlights several critical optimization points: macrophage differentiation quality, antibiotic selection, polybrene tolerance, VPX-VLP functionality, lentiviral MOI determination, Cas9 expression, knockout validation, and guide RNA distribution. Together, these elements support more reliable CRISPR screening in a cell type that is biologically important but technically challenging.
For inflammation, infection, cancer, neurodegeneration, and immune regulation research, optimized CRISPR-Cas9 knockout in iPSC-derived macrophages offers a route toward discovering key regulators of macrophage function. The strongest results will come from workflows that combine careful technical optimization with rigorous biological validation.
This article is based on the scientific content provided from:
Navarro-Guerrero, E., Baronio, R., Tay, C., Knight, J. C., & Ebner, D. V. “Optimized protocol for CRISPR knockout of human iPSC-derived macrophages.” STAR Protocols, Volume 5, Issue 1, 2024.