- IGram-negative bacteria are the pink or red-colored bacteria when observed under a microscope after Gram’s differential staining technique.
- They do not retain the crystal violet stain due to its thinner layer cell wall that completely removes the dye by the solvent used and the safranin stains the cell of the gram-negative bacteria in pink or red color.
- Bacteria can be found everywhere that may support our life or may even harm us. Some gram-negative pathogenic bacteria such as Escherichia coli, Pseudomonas aeruginosa, Chlamydia trachomatis, Yersinia pestis causes serious health diseases on living organisms.
- Some of the bacterial infections caused by gram-negative cells are Cholera, Brucellosis, Typhoid fever, Campylobacter infection, E. coli infection, Plague, and Shigellosis.
- Some of the antibiotics used for gram-negative bacteria are cephalosporins (ceftriaxone-cefotaxime, ceftazidime), fluoroquinolones (ciprofloxacin, levofloxacin), aminoglycosides (gentamicin, amikacin), imipenem, penicillin (amoxicillin-clavulanic acid, piperacillin-tazobactam), and trimethoprim-sulfamethoxazole.
- Gram-negative bacteria are more antibiotic-resistant than gram-positive bacteria due to its presence of a largely impermeable cell wall that alters during the mutation and changing the hydrophobic properties of its outer membrane which lacks in gram-positive bacteria.
- To identify the bacterial infection and drug discovery, species of gram-negative bacteria must be identified using morphological as well as microbiological tests such as morphological observation of colonies, Gram staining test, and several biochemical tests following the Bergey’s manual.
Identify gram negative bacteria precisely with biochemical tests. Learn how to perform the tests here and identify the organism here.
Morphological test
The colony morphology of organisms is observed. Colony shape, size (in mm), color, consistency, elevation, opacity, and margin are observed and noted down for further identification.
Gram staining for identifying gram negative bacteria
The Gram staining is the beginning test in an identification procedure in bacterial classification. Gram staining is a differential staining technique that differentiates gram-positive and gram-negative bacteria from a cultured colony. The pink or red color observed under the microscope after performing Gram staining confirms the gram-negative bacteria whereas gram positive appears purple in color.
The difference in color between these two bacteria is because of their different composition of cell wall layer. Similarly, the mechanism of the Gram stain also differs in the permeability of the bacterial cell wall in the decolourizing step.
The cell wall of gram-positive bacteria contains a thick peptidoglycan layer with low lipid content and is more sensitive to alcohol used in the decolourizing step which closes up the pores of the wall making the primary dye crystal violet unable to escape, and thus the primary purple color is retained. Gram-negative cell wall contains higher lipid content which gets dissolved during decolorization which opens up the pore size and the primary dye leach out and takes up the counterstain and appears pink or red in color.
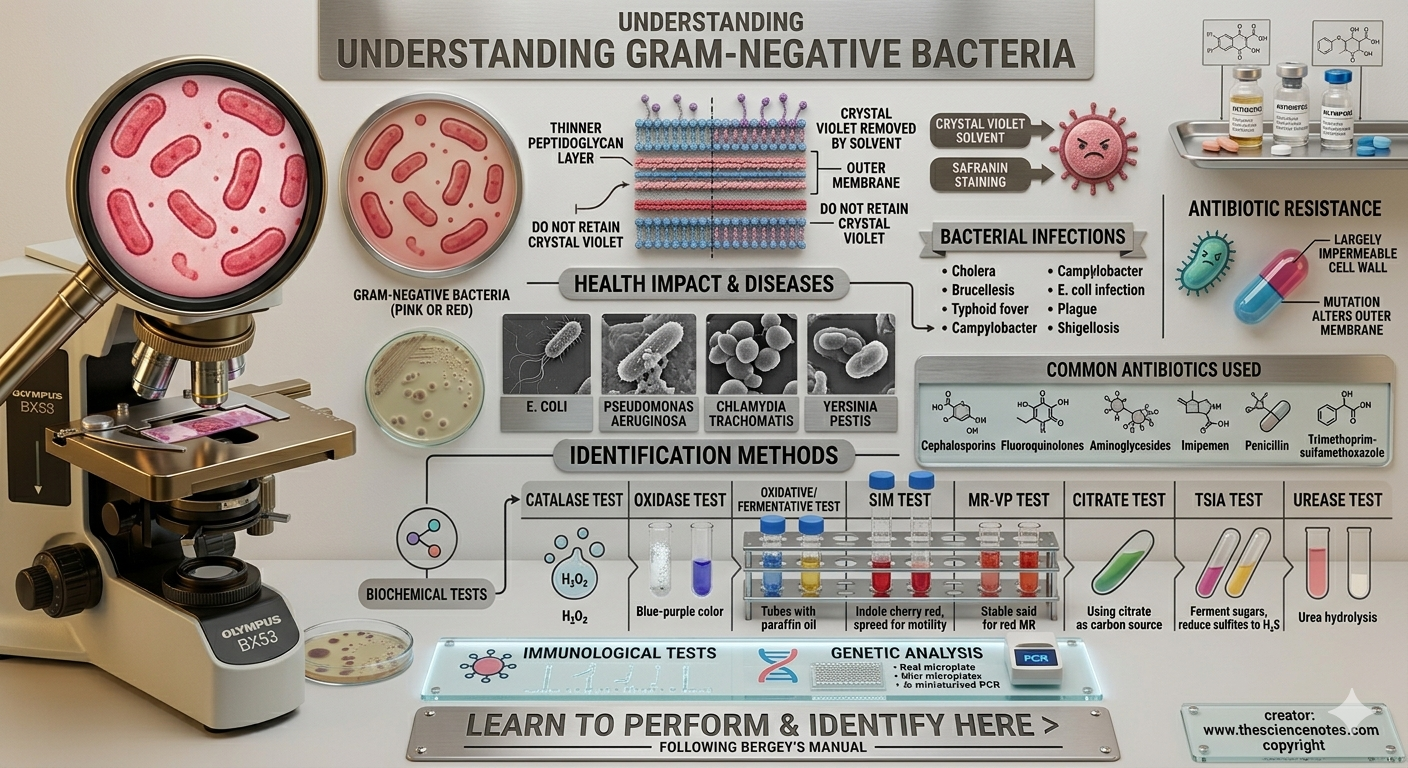
Biochemical tests for the identification of gram negative bacterial species
A biochemical test determines the organism’s enzymatic activity on specific substances used during the test which helps in the identification of the bacterial species by their action of enzymes. Immunological tests and genetic analysis are also required in addition to identifying.
Some of the specific biochemical tests done to identify gram negative organisms are:
- Catalase
- Oxidase
- Oxidative/Fermentative
- SIM
- MR-VP
- Citrate
- TSIA
- Urease
Catalase test
- Catalase enzyme is produced by most of the gram-negative organisms which in reaction with 3% H2O2 gives an effervescence of gas bubbles.
- Catalase positive gram-negative species are:
Escherichia coli, Proteus mirabilis, Klebsiella pneumoniae, Klebsiella oxytoca, Shigella dysentry, Shigella flexneri, Salmonella Paratyphi, Salmonella Typhirium, Pseudomonas aeruginosa, Seratia marcescens, Enterobacter aerogens, Citrobacter fruendii.
Oxidase test
- Oxidase producing gram-negative bacteria develops dark blue-purple color when rubbed with a filter paper soaked in the tetramethyl-p-phenylenediamine reagent.
- Oxidase positive gram-negative species are:
Pseudomonas aeruginosa, Helicobacter pylori, Vibrio cholerae, Campylobacter jejuni.
Oxidative/Fermentative test
- Oxidative organism utilizes the substrate in aerobic condition whereas fermentative organism utilizes in anaerobic condition.
- To maintain an anaerobic condition, one tube is sealed with paraffin oil on the top layer.
- Oxidative gram-negative species are:
Pseudomonas aeruginosa, P. vesicularis, P. alcaligenes, A. faecalis, A. denitrificans.
- Fermentative gram negative species are:
E. coli, Proteus mirabilis, K. pneumoniae, K. oxytoca, S, dysentry, E. aerogens.
SIM test
- Sulfur Indole Motility test is for sulfide production and motility which is indicated by the formation of cherry red color in indole test and bacterial spread in media indicates motile organism.
- Indole positive organisms are:
E. coli, K. oxytoca, S. dysentry
- Motile organisms are:
E. coli, P. mirabilis, S. dysentry, S. Paratyphi, S. Typhirium, C. fruendii.
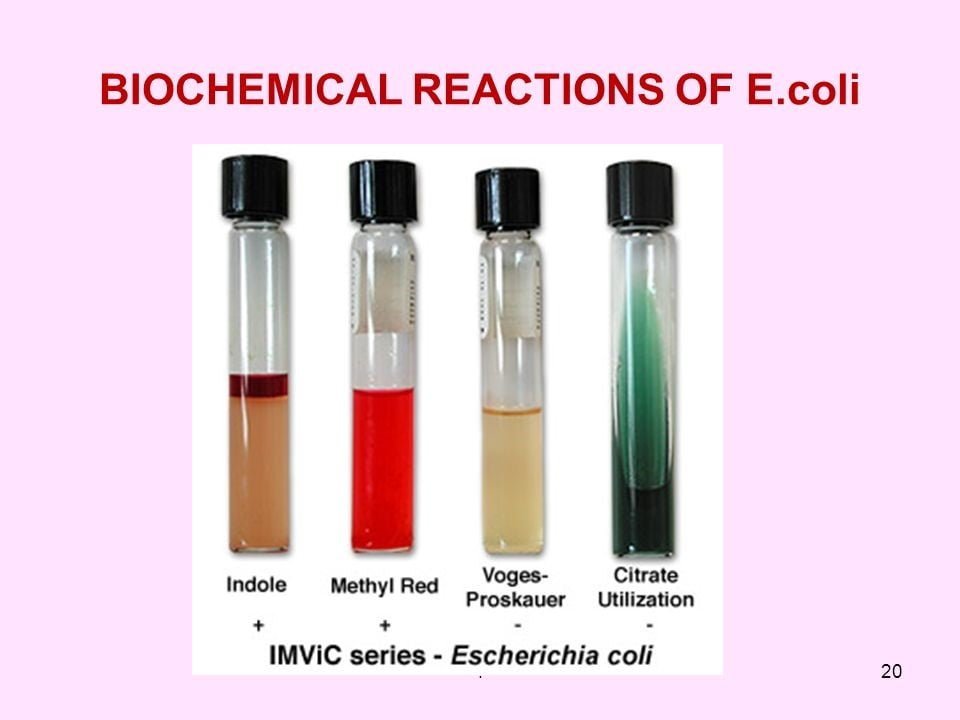

MR-VP test
- Methyl red-Voges Proskauer test is for the detection of organisms able to maintain stable acid which gives red color in case of the positive result.
- MR positive organisms are:
E. coli, S. dysentry, S flexneri, E. aerogens, C. fruendii
- VP positive organisms are:
K. pneumoniae, K. oxytoca, S. marcescens, E. aerogens
Citrate test
- The citrate test detects the organism using citrate as a sole source of carbon during metabolism.
- Citrate positive organisms are:
P. mirabilis, K. pneumoniae, K. oxytoca, P. aeruginosa, C. fruendii
TSIA (Triple Sugar Iron Agar) Test
- This test is for the organisms ability to ferment sugars and reduce sulfites to sulfides.
- Acid and alkali producing organisms are:
E. coli, K. pneumoniae, S. marcescens, E. aerogens
- H2S producing organisms are:
P. mirabilis, S. Typhirium, C. fruendii
Urease test
- Urea hydrolysis test detects the organism producing urease enzyme.
- Urease positive gram negative organisms are:
K. pneumoniae, K. oxytoca, P. mirabilis
References:
- Manandhar.S,Sharma.S,(2013).Practical Approach to Microbiology,National Book Center,2nd edition (Pageno.65-81).