The trp Operon Transcription Attenuation mechanism represents one of the most elegant examples of genetic economy in bacteria. While many operons rely solely on a simple “on/off” switch at the initiation of transcription, Escherichia coli utilizes a secondary layer of control to fine-tune the biosynthesis of tryptophan. This process, known as trp Operon Transcription Attenuation, allows the cell to sense the immediate availability of charged tRNA molecules and halt mRNA synthesis before the structural genes are even reached. By coupling transcription directly to translation, the cell ensures that energy is never wasted on redundant metabolic pathways.
1. The Dual-Level Control of the trp Operon
The E. coli trp operon is a repressible system regulated at two distinct levels. The first level involves the Trp Repressor, a homodimer that blocks the initiation of transcription when tryptophan levels are high. However, once repression is lifted and RNA polymerase begins its journey, the second regulatory process—trp Operon Transcription Attenuation—takes over to provide high-resolution control based on the metabolic state of the cell.
The Structural Genes
The operon contains five structural genes (trpE, trpD, trpC, trpB, and trpA) that catalyze the formation of tryptophan from chorismate. Together, these enzymes are the workhorses of tryptophan biosynthesis, but their production is strictly governed by the trp Operon Transcription Attenuation signal found in the 5′ leader sequence.
2. Molecular Mechanics of trp Operon Transcription Attenuation
The heart of trp Operon Transcription Attenuation lies within a 161-nucleotide leader RNA sequence (trpL). This sequence contains a short “leader peptide” and four regions that can form alternative secondary structures through intramolecular base pairing.
The Four Regulatory Regions
Region 1: Contains two consecutive Trp codons. This acts as the “sensor” for tryptophan availability.
Region 2: Can pair with Region 3 to form an antiterminator.
Region 3: Can pair with either Region 2 (antitermination) or Region 4 (termination).
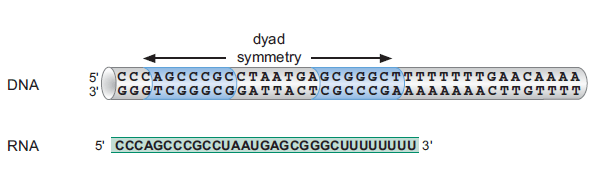
Region 4: Pairs with Region 3 to form the attenuator loop, a rho-independent terminator followed by a Poly-U track.
3. High Tryptophan Levels and the Termination Signal
In a high-tryptophan environment, the trp Operon Transcription Attenuation mechanism favors the termination of transcription. Because charged $tRNA^{Trp}$ is abundant, the ribosome translating the leader peptide does not stall at the Trp codons in Region 1. Instead, it moves quickly into Region 2.
When the ribosome covers Region 2, it prevents it from pairing with Region 3. Consequently, as RNA polymerase transcribes Region 4, the 3:4 termination loop forms. This structure signals the RNA polymerase to fall off the DNA template, effectively attenuating transcription before the structural genes are transcribed.
4. Low Tryptophan Levels and the Antitermination Response
When the cellular supply of tryptophan is low, the trp Operon Transcription Attenuation pathway switches to an “express” mode. In this scenario, the ribosome begins translating the leader peptide but stalls at the two consecutive Trp codons in Region 1 because it cannot find enough charged $tRNA^{Trp}$.
This stalling leaves Region 2 exposed. As RNA polymerase transcribes Region 3, Region 2 and Region 3 immediately base pair to form the 2:3 antitermination loop. Because Region 3 is now occupied, it cannot pair with Region 4. Without the 3:4 terminator loop, RNA polymerase continues through the attenuator and transcribes the structural genes (trpEDCBA), allowing the cell to synthesize the tryptophan it lacks.
5. Evolutionary Conservation: Beyond the trp Operon
The principles of trp Operon Transcription Attenuation are not restricted to tryptophan. Many other amino acid biosynthetic operons in E. coli utilize similar “leader peptide” sensors:
The his Operon: Features seven consecutive histidine codons in its leader sequence.
The leu Operon: Features four consecutive leucine codons.
Unlike the trp system, which uses both a repressor and an attenuator, the his and leu operons rely almost exclusively on transcription attenuation to govern their expression. This highlights the evolutionary success of RNA secondary structures as metabolic sensors.
6. Summary of trp Operon Transcription Attenuation Logic
| Tryptophan Level | Ribosome Status | RNA Structure Formed | Final Outcome |
| High | Translates through Reg 1 | 3:4 Loop (Terminator) | Transcription Terminated |
| Low | Stalls at Reg 1 (Trp codons) | 2:3 Loop (Antiterminator) | Structural Genes Transcribed |
7. Conclusion: The Precision of RNA-Based Sensing
The trp Operon Transcription Attenuation system is a masterpiece of molecular engineering. It demonstrates that gene regulation is a multi-layered process where the physical speed of a ribosome can dictate the folding of an RNA molecule, which in turn determines the fate of a whole metabolic pathway. Understanding trp Operon Transcription Attenuation provides deep insights into how prokaryotes maintain homeostasis in a fluctuating environment.
1. The Architecture of the trp Operon
The trp operon is a repressible operon responsible for the biosynthesis of the amino acid tryptophan from chorismate. It consists of five structural genes that encode the enzymes necessary for this metabolic pathway:
trpE & trpD: Anthranilate Synthase Components I and II.
trpC: N-(5’-Phosphoribosyl)-Anthranilate Isomerase and Indole-3-Glycerol Phosphate Synthase.
trpB & trpA: Tryptophan Synthase $\beta$ and $\alpha$ subunits.
Beyond these structural genes lies the regulatory heart of the system: the promoter, the operator, and the leader sequence (trpL), which contains the attenuator region.

STRUCTURE OF TRP OPERON
2. Level One: The Trp Repressor (Initiation Control)
The first line of defense against wasting resources is the Trp Repressor. This homodimeric protein contains a helix-turn-helix motif designed to bind to the operator sequence, which overlaps with the promoter.
High Tryptophan Levels: Tryptophan acts as a corepressor. It binds to the repressor, causing a conformational change that allows the repressor to sit firmly on the operator. This physically blocks RNA Polymerase from initiating transcription.
Low Tryptophan Levels: The repressor loses its tryptophan molecules and falls off the DNA. RNA Polymerase is then free to bind to the promoter and begin transcribi
ng the mRNA.
3. Level Two: The Mechanism of Attenuation
Even after repression is lifted and transcription begins, the cell employs a “fine-tuning” mechanism called attenuation. Attenuation regulates the operon by controlling the termination of transcription at a site located before the first structural gene (trpE).
The Leader Sequence and the 4 Regions
Analysis of the 161-nucleotide leader RNA reveals four distinct regions capable of forming alternative secondary structures through intramolecular base pairing:
Region 1: Contains the coding sequence for a short leader peptide, including two consecutive tryptophan codons.
Region 2: Can base pair with Region 1 or Region 3.
Region 3: Can base pair with Region 2 or Region 4.
Region 4: When paired with Region 3, forms a G-C rich stem-loop followed by a poly-U sequence—a classic rho-independent transcription terminator.
4. The Coupling of Transcription and Translation
In bacteria, transcription and translation occur in the same compartment simultaneously. This “coupling” is essential for attenuation. As RNA Polymerase transcribes the leader sequence, a ribosome immediately attaches to the nascent mRNA to translate the leader peptide.
Scenario A: High Tryptophan Levels (Termination)
When tryptophan is abundant, charged $tRNA^{Trp}$ is readily available. The ribosome translates the leader peptide rapidly and moves into Region 2.
Because the ribosome is covering Region 2, it cannot pair with Region 3.
As RNA Polymerase transcribes Region 3 and Region 4, they are free to base pair with each other.
This forms the 3:4 termination loop (the attenuator).
The RNA Polymerase falls off, and the structural genes are not transcribed.
Scenario B: Low Tryptophan Levels (Antitermination)
When tryptophan is scarce, charged $tRNA^{Trp}$ is unavailable. The ribosome reaches the two Trp codons in Region 1 and stalls, waiting for a $tRNA$ that isn’t coming.
The stalled ribosome leaves Region 2 exposed.
As soon as RNA Polymerase transcribes Region 3, Region 2 and Region 3 immediately base pair.
The formation of the 2:3 antitermination loop prevents the formation of the 3:4 loop.
With no termination signal in its way, RNA Polymerase continues past the attenuator to transcribe the structural genes, allowing the cell to make its own tryptophan.

5. Attenuation in Other Operons
The trp operon is the most famous example, but it is not unique. Many amino
acid biosynthetic pathways rely on this “sensing” of $tRNA$ levels.
his Operon: The leader sequence contains seven histidine codons in a row.
leu Operon: The leader sequence contains four leucine codons.
Interestingly, while the trp operon uses both repression and attenuation, the his and leu operons in E. coli rely almost entirely on attenuation for their regulation.
6. Clinical and Research Significance
Understanding secondary RNA structures and attenuation provides novel avenues for synthetic biology and drug development. By manipulating these leader sequences, researchers can create “riboswitches” that respond to specific molecular signals, a technique used in developing biosensors and advanced metabolic engineering.
Furthermore, because attenuation relies on the coupling of transcription and
translation—a process unique to prokaryotes—it represents a potential target for narrow-spectrum antibiotics that would not interfere with eukaryotic (human) gene expression.
7. Summary Table: Trp Operon Logic
| Tryptophan Level | Ribosome Action | RNA Structure | Outcome |
| High | Moves to Region 2 | 3:4 Loop (Terminator) | Transcription stops at Attenuator |
| Low | Stalls at Region 1 | 2:3 Loop (Antiterminator) | Transcription of structural genes |
Conclusion
The trp operon stands as a masterclass in biological economy. By using the very process of translation to regulate transcription, bacteria achieve a level of sensitivity and responsiveness that ensures their survival in ever-changing environments. It reminds us that gene regulation is not just about “on” and “off” switches, but a series of fluid, structural checkpoints.
References
Yanofsky, C. (1981). “Attenuation in the control of expression of bacterial operons.” Nature.
Watson, J. D., et al. (2013). “Molecular Biology of the Gene.”
NCBI Bookshelf – The trp Operon: A Model of Gene Regulation.